Search by Genic region
| geneid | 4184 |
|---|---|
| ensemblid | ENSG00000163206.6 |
| hgncid | 6962 |
| symbol | SMCP |
| name | sperm mitochondria associated cysteine rich protein |
| refseq_nuc | NM_030663.3 |
| refseq_prot | NP_109588.2 |
| ensembl_nuc | ENST00000368765.4 |
| ensembl_prot | ENSP00000357754.3 |
| mane_status | MANE Select |
| chr | chr1 |
| start | 152878322 |
| end | 152885047 |
| strand | + |
| ver | v1.2 |
| region | chr1:152878322-152885047 |
| region5000 | chr1:152873322-152890047 |
| regionname0 | SMCP_chr1_152878322_152885047 |
| regionname5000 | SMCP_chr1_152873322_152890047 |
| ahapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
alen | total | AFR | AMR | EAS | EUR | SAS | JPT | regionname | genename | aa | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001 | 1/1 | 116 | 395 | 78 | 69 | 198 | 10 | 38 | 152 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0002 | 0/0 | 116 | 22 | 21 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0003 | 0/0 | 116 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| chapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
clen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | cseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| c0001 | 1/1 | 351 | 395 | 78 | 69 | 198 | 10 | 38 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| c0002 | 0/0 | 351 | 21 | 20 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| c0003 | 0/0 | 351 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| c0004 | 0/0 | 351 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| thapid | grch38chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
tlen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | tseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| t0001 | 1/1 | 420 | 272 | 71 | 59 | 102 | 10 | 28 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| t0002 | 0/0 | 420 | 115 | 0 | 10 | 95 | 0 | 10 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| t0003 | 0/0 | 420 | 20 | 19 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| t0004 | 0/0 | 420 | 7 | 7 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| t0005 | 0/0 | 420 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| t0006 | 0/0 | 420 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| t0007 | 0/0 | 420 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| ghapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
total | AFR | AMR | EAS | EUR | SAS | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| g0001 | 0/0 | 165 | 10 | 49 | 79 | 6 | 21 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0002 | 0/0 | 86 | 0 | 8 | 71 | 0 | 7 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0003 | 0/0 | 18 | 6 | 0 | 11 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0004 | 0/0 | 13 | 0 | 1 | 10 | 0 | 2 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0005 | 0/0 | 12 | 11 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0006 | 0/0 | 11 | 2 | 5 | 1 | 3 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0007 | 0/0 | 10 | 10 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0008 | 0/0 | 8 | 8 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0009 | 0/0 | 7 | 7 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0010 | 0/0 | 6 | 5 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0011 | 0/0 | 6 | 6 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0012 | 0/0 | 5 | 5 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0013 | 0/0 | 5 | 5 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0014 | 0/0 | 4 | 4 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0015 | 0/0 | 4 | 4 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0016 | 0/0 | 4 | 0 | 1 | 3 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0017 | 0/0 | 3 | 0 | 0 | 1 | 0 | 2 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0018 | 0/0 | 2 | 0 | 2 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0019 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0020 | 0/0 | 2 | 1 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0021 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0022 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0023 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0024 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0025 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0026 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0027 | 1/0 | 1 | 0 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0028 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0029 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0030 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0031 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0032 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0033 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0034 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0035 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0036 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0037 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0038 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0039 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0040 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0041 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0042 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0043 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0044 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0045 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0046 | 0/1 | 1 | 0 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0047 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0048 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0049 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0050 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0051 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0052 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0053 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0054 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0055 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0056 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0057 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0058 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0059 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0060 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0061 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| g0062 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| achapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
clen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | cseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001 | 1/1 | 351 | 395 | 78 | 69 | 198 | 10 | 38 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0002c0002 | 0/0 | 351 | 21 | 20 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0002c0004 | 0/0 | 351 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0003c0003 | 0/0 | 351 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| acthapid | grch38chm13v2 | tlen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | tseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001t0001 | 1/1 | 770 | 271 | 70 | 59 | 102 | 10 | 28 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0001c0001t0002 | 0/0 | 770 | 115 | 0 | 10 | 95 | 0 | 10 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0001c0001t0004 | 0/0 | 770 | 7 | 7 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0001c0001t0006 | 0/0 | 770 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0001c0001t0007 | 0/0 | 770 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0002c0002t0003 | 0/0 | 770 | 20 | 19 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0002c0002t0005 | 0/0 | 770 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0002c0004t0005 | 0/0 | 770 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| a0003c0003t0001 | 0/0 | 770 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | copy fasta | chr1 | 152873322 | 152890047 |
| actghapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
total | AFR | AMR | EAS | EUR | SAS | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001t0001g0001 | 0/0 | 165 | 10 | 49 | 79 | 6 | 21 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0003 | 0/0 | 18 | 6 | 0 | 11 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0006 | 0/0 | 10 | 2 | 5 | 0 | 3 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0007 | 0/0 | 10 | 10 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0008 | 0/0 | 8 | 8 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0010 | 0/0 | 6 | 5 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0011 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0012 | 0/0 | 4 | 4 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0013 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0014 | 0/0 | 4 | 4 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0015 | 0/0 | 4 | 4 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0017 | 0/0 | 3 | 0 | 0 | 1 | 0 | 2 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0018 | 0/0 | 2 | 0 | 2 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0019 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0020 | 0/0 | 2 | 1 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0021 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0022 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0024 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0025 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0026 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0027 | 1/0 | 1 | 0 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0028 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0029 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0030 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0031 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0032 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0033 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0034 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0035 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0036 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0037 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0038 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0039 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0040 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0044 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0045 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0046 | 0/1 | 1 | 0 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0048 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0049 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0001g0051 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0002 | 0/0 | 86 | 0 | 8 | 71 | 0 | 7 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0004 | 0/0 | 13 | 0 | 1 | 10 | 0 | 2 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0016 | 0/0 | 4 | 0 | 1 | 3 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0023 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0052 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0053 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0054 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0055 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0056 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0057 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0058 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0059 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0060 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0002g0061 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0004g0009 | 0/0 | 7 | 7 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0006g0006 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0001c0001t0007g0012 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0005 | 0/0 | 12 | 11 | 1 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0011 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0013 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0041 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0042 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0043 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0003g0050 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0002t0005g0011 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0002c0004t0005g0062 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| a0003c0003t0001g0047 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| sampleid | ID haplotypeid
|
ahapid | chapid | thapid | ghapid | gpopname | popname | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| HG00140 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | GBR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00140 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | GBR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00280 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | FIN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00280 | hp2 | a0001 | c0001 | t0001 | g0051 | EUR | FIN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00408 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00408 | hp2 | a0001 | c0001 | t0002 | g0052 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00423 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00423 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00438 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00438 | hp2 | a0001 | c0001 | t0001 | g0038 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00544 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00544 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00558 | hp1 | a0001 | c0001 | t0001 | g0019 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00558 | hp2 | a0001 | c0001 | t0002 | g0016 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00597 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00597 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00609 | hp1 | a0001 | c0001 | t0001 | g0025 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00609 | hp2 | a0001 | c0001 | t0001 | g0003 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00621 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00621 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00642 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00642 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00673 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00673 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | CHS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00733 | hp1 | a0001 | c0001 | t0001 | g0006 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00733 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00735 | hp1 | a0001 | c0001 | t0001 | g0037 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00735 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00738 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00738 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00741 | hp1 | a0001 | c0001 | t0001 | g0006 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG00741 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01069 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01069 | hp2 | a0001 | c0001 | t0002 | g0016 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01071 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01071 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01074 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01074 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01099 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01099 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01106 | hp1 | a0001 | c0001 | t0002 | g0004 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01106 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01109 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01109 | hp2 | a0001 | c0001 | t0001 | g0006 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01167 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01167 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01168 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01168 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01175 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01175 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01243 | hp1 | a0001 | c0001 | t0001 | g0026 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01243 | hp2 | a0001 | c0001 | t0001 | g0018 | AMR | PUR | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01255 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01255 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01256 | hp1 | a0001 | c0001 | t0002 | g0002 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01256 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01257 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01257 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01258 | hp1 | a0001 | c0001 | t0002 | g0002 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01258 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01346 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01346 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01358 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01358 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01361 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01361 | hp2 | a0001 | c0001 | t0002 | g0002 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01433 | hp1 | a0002 | c0002 | t0003 | g0005 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01433 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01496 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01496 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01515 | hp1 | a0001 | c0001 | t0001 | g0006 | EUR | IBS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01515 | hp2 | a0001 | c0001 | t0001 | g0006 | EUR | IBS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01516 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | IBS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01516 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | IBS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01517 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | IBS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01517 | hp2 | a0001 | c0001 | t0001 | g0006 | EUR | IBS | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01884 | hp1 | a0001 | c0001 | t0001 | g0006 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01884 | hp2 | a0002 | c0002 | t0003 | g0013 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01891 | hp1 | a0001 | c0001 | t0001 | g0021 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01891 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01934 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01934 | hp2 | a0001 | c0001 | t0001 | g0018 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01943 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01943 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01952 | hp1 | a0001 | c0001 | t0002 | g0002 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01952 | hp2 | a0001 | c0001 | t0002 | g0002 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01975 | hp1 | a0001 | c0001 | t0001 | g0020 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01975 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01978 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01978 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01981 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01981 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02004 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02004 | hp2 | a0001 | c0001 | t0001 | g0006 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02015 | hp1 | a0001 | c0001 | t0002 | g0016 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02015 | hp2 | a0001 | c0001 | t0006 | g0006 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02027 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02027 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02040 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02040 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02055 | hp1 | a0001 | c0001 | t0001 | g0012 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02055 | hp2 | a0001 | c0001 | t0001 | g0013 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02056 | hp1 | a0001 | c0001 | t0002 | g0004 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02056 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02071 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02071 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02074 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02074 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02080 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02080 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02083 | hp1 | a0001 | c0001 | t0001 | g0003 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02083 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02129 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02129 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02132 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02132 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02135 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02135 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | KHV | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02145 | hp1 | a0001 | c0001 | t0001 | g0044 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02145 | hp2 | a0002 | c0002 | t0003 | g0013 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02148 | hp1 | a0001 | c0001 | t0002 | g0002 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02148 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02165 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CDX | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02165 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | CDX | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02257 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02257 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02258 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02258 | hp2 | a0001 | c0001 | t0001 | g0012 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02280 | hp1 | a0002 | c0002 | t0003 | g0011 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02280 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02293 | hp1 | a0001 | c0001 | t0002 | g0002 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02293 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02300 | hp1 | a0001 | c0001 | t0001 | g0006 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02300 | hp2 | a0001 | c0001 | t0002 | g0002 | AMR | PEL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02451 | hp1 | a0001 | c0001 | t0001 | g0007 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02451 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02572 | hp1 | a0002 | c0002 | t0003 | g0005 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02572 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02602 | hp1 | a0001 | c0001 | t0002 | g0002 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02602 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02615 | hp1 | a0002 | c0002 | t0003 | g0011 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02615 | hp2 | a0001 | c0001 | t0001 | g0010 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02622 | hp1 | a0001 | c0001 | t0001 | g0045 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02622 | hp2 | a0001 | c0001 | t0004 | g0009 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02630 | hp1 | a0003 | c0003 | t0001 | g0047 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02630 | hp2 | a0001 | c0001 | t0004 | g0009 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02647 | hp1 | a0001 | c0001 | t0001 | g0020 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02647 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02683 | hp1 | a0001 | c0001 | t0002 | g0002 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02683 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02698 | hp1 | a0001 | c0001 | t0001 | g0024 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02698 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02717 | hp1 | a0001 | c0001 | t0001 | g0015 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02717 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02723 | hp1 | a0001 | c0001 | t0001 | g0010 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02723 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02735 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02735 | hp2 | a0001 | c0001 | t0001 | g0039 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02738 | hp1 | a0001 | c0001 | t0002 | g0004 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02738 | hp2 | a0001 | c0001 | t0001 | g0017 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02809 | hp1 | a0001 | c0001 | t0004 | g0009 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02809 | hp2 | a0002 | c0002 | t0005 | g0011 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02818 | hp1 | a0001 | c0001 | t0001 | g0015 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02818 | hp2 | a0001 | c0001 | t0001 | g0028 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02886 | hp1 | a0002 | c0002 | t0003 | g0005 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02886 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02895 | hp1 | a0001 | c0001 | t0001 | g0014 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02895 | hp2 | a0001 | c0001 | t0001 | g0012 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02896 | hp1 | a0002 | c0002 | t0003 | g0005 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02896 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02897 | hp1 | a0001 | c0001 | t0001 | g0012 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02897 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02922 | hp1 | a0001 | c0001 | t0001 | g0013 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02922 | hp2 | a0002 | c0002 | t0003 | g0050 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02965 | hp1 | a0001 | c0001 | t0001 | g0040 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02965 | hp2 | a0001 | c0001 | t0001 | g0022 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02970 | hp1 | a0001 | c0001 | t0001 | g0008 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02970 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02976 | hp1 | a0001 | c0001 | t0001 | g0011 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02976 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03017 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03017 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03041 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03041 | hp2 | a0001 | c0001 | t0004 | g0009 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03098 | hp1 | a0001 | c0001 | t0001 | g0007 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03098 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03130 | hp1 | a0001 | c0001 | t0001 | g0007 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03130 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03139 | hp1 | a0001 | c0001 | t0001 | g0014 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03139 | hp2 | a0001 | c0001 | t0004 | g0009 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03195 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03195 | hp2 | a0001 | c0001 | t0004 | g0009 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03209 | hp1 | a0001 | c0001 | t0001 | g0014 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03209 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03225 | hp1 | a0001 | c0001 | t0001 | g0014 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03225 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03239 | hp1 | a0001 | c0001 | t0001 | g0032 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03239 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03453 | hp1 | a0001 | c0001 | t0001 | g0010 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03453 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03486 | hp1 | a0001 | c0001 | t0007 | g0012 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03486 | hp2 | a0001 | c0001 | t0001 | g0048 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03491 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03491 | hp2 | a0001 | c0001 | t0002 | g0002 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03492 | hp1 | a0001 | c0001 | t0002 | g0004 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03492 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03516 | hp1 | a0002 | c0004 | t0005 | g0062 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03516 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | ESN | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03540 | hp1 | a0002 | c0002 | t0003 | g0043 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03540 | hp2 | a0001 | c0001 | t0004 | g0009 | AFR | GWD | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03579 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03579 | hp2 | a0002 | c0002 | t0003 | g0041 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03654 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03654 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03669 | hp1 | a0001 | c0001 | t0001 | g0003 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03669 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03688 | hp1 | a0001 | c0001 | t0002 | g0002 | SAS | STU | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03688 | hp2 | a0001 | c0001 | t0001 | g0017 | SAS | STU | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03704 | hp1 | a0001 | c0001 | t0002 | g0002 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03704 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03710 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03710 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03834 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03834 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03927 | hp1 | a0001 | c0001 | t0001 | g0010 | SAS | BEB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03927 | hp2 | a0001 | c0001 | t0002 | g0002 | SAS | BEB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG04184 | hp1 | a0001 | c0001 | t0002 | g0055 | SAS | BEB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG04184 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG04228 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG04228 | hp2 | a0001 | c0001 | t0002 | g0002 | SAS | STU | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18522 | hp1 | a0001 | c0001 | t0001 | g0011 | AFR | YRI | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18522 | hp2 | a0001 | c0001 | t0001 | g0010 | AFR | YRI | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18612 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18612 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18747 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CHB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18747 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | CHB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18906 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | YRI | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18906 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | YRI | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18940 | hp1 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18940 | hp2 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18941 | hp1 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18941 | hp2 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18942 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18942 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18943 | hp1 | a0001 | c0001 | t0001 | g0031 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18943 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18945 | hp1 | a0001 | c0001 | t0002 | g0023 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18945 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18946 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18946 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18948 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18948 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18949 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18949 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18950 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18950 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18951 | hp1 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18951 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18952 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18952 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18953 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18953 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18954 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18954 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18957 | hp1 | a0001 | c0001 | t0002 | g0016 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18957 | hp2 | a0001 | c0001 | t0001 | g0036 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18959 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18959 | hp2 | a0001 | c0001 | t0002 | g0058 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18960 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18960 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18963 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18963 | hp2 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18966 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18966 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18967 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18967 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18968 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18968 | hp2 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18969 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18969 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18970 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18970 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18971 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18971 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18973 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18973 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18975 | hp1 | a0001 | c0001 | t0002 | g0061 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18975 | hp2 | a0001 | c0001 | t0002 | g0056 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18982 | hp1 | a0001 | c0001 | t0001 | g0034 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18982 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18983 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18983 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18984 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18984 | hp2 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18985 | hp1 | a0001 | c0001 | t0002 | g0057 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18985 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18989 | hp1 | a0001 | c0001 | t0001 | g0017 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18989 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18990 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18990 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18991 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18991 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18993 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18993 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18994 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18994 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18995 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18995 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18997 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18997 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18998 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18998 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18999 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18999 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19000 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19000 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19001 | hp1 | a0001 | c0001 | t0002 | g0054 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19001 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19002 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19002 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19003 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19003 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19005 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19005 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19006 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19006 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19007 | hp1 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19007 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19010 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19010 | hp2 | a0001 | c0001 | t0001 | g0033 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19011 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19011 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19012 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19012 | hp2 | a0001 | c0001 | t0002 | g0053 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19030 | hp1 | a0001 | c0001 | t0001 | g0013 | AFR | LWK | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19030 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | LWK | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19043 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | LWK | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19043 | hp2 | a0001 | c0001 | t0001 | g0049 | AFR | LWK | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19056 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19056 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19057 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19057 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19058 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19058 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19062 | hp1 | a0001 | c0001 | t0002 | g0059 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19062 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19063 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19063 | hp2 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19064 | hp1 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19064 | hp2 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19065 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19065 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19067 | hp1 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19067 | hp2 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19068 | hp1 | a0001 | c0001 | t0001 | g0030 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19068 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19070 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19070 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19072 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19072 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19075 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19075 | hp2 | a0001 | c0001 | t0001 | g0035 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19077 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19077 | hp2 | a0001 | c0001 | t0002 | g0060 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19078 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19078 | hp2 | a0001 | c0001 | t0001 | g0019 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19079 | hp1 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19079 | hp2 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19080 | hp1 | a0001 | c0001 | t0002 | g0023 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19080 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19081 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19081 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19082 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19082 | hp2 | a0001 | c0001 | t0002 | g0004 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19083 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19083 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19084 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19084 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19085 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19085 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19086 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19086 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19087 | hp1 | a0001 | c0001 | t0001 | g0003 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19087 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19088 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19088 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19089 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19089 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19090 | hp1 | a0001 | c0001 | t0001 | g0029 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19090 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19091 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19091 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19240 | hp1 | a0001 | c0001 | t0001 | g0015 | AFR | YRI | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA19240 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | YRI | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA20129 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | ASW | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA20129 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | ASW | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA20905 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | GIH | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA20905 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | GIH | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01123 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG01123 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02109 | hp1 | a0001 | c0001 | t0001 | g0021 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02109 | hp2 | a0002 | c0002 | t0003 | g0042 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02486 | hp1 | a0001 | c0001 | t0001 | g0008 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02486 | hp2 | a0001 | c0001 | t0001 | g0011 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02559 | hp1 | a0001 | c0001 | t0001 | g0006 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG02559 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | ACB | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03471 | hp1 | a0002 | c0002 | t0003 | g0005 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG03471 | hp2 | a0002 | c0002 | t0003 | g0005 | AFR | MSL | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG06807 | hp1 | a0001 | c0001 | t0001 | g0010 | AFR | USA | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| HG06807 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | USA | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18955 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA18955 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA20300 | hp1 | a0001 | c0001 | t0001 | g0022 | AFR | USA | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA20300 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | USA | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA21309 | hp1 | a0001 | c0001 | t0001 | g0015 | AFR | LWK | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| NA21309 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | LWK | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| homoSapiens_chm13v2 | hp1 | a0001 | c0001 | t0001 | g0046 | REF | REF | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| homoSapiens_grch38 | hp1 | a0001 | c0001 | t0001 | g0027 | REF | REF | SMCP_chr1_152873322_152890047 | SMCP | chr1 | 152873322 | 152890047 |
| chr:pos | ref | alt | # # of ahapid:amino-acid(protein) level |
ahapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:152884488
|
G | T | 1 | a0003 | 1 | HG02630.hp1 | missense_variant | MODERATE | c.66G>T | p.Gln22His | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 211/770 | 66/351 | 22/116 | chr1 | 152884488 | ||
| chr1:152884519
|
C | G | 1 | a0002 | 22 | HG01433.hp1 HG01884.hp2 HG02109.hp2 others(19): Show |
missense_variant | MODERATE | c.97C>G | p.Gln33Glu | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 242/770 | 97/351 | 33/116 | chr1 | 152884519 |
| chr:pos | ref | alt | # # of chapid |
chapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:152884461
|
A | G | 1 | a0002c0004 | 1 | HG03516.hp1 | synonymous_variant | LOW | c.39A>G | p.Ala13Ala | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 184/770 | 39/351 | 13/116 | chr1 | 152884461 | ||
| chr1:152884626
|
G | A | 2 | a0002c0002a0002c0004 | 22 | HG01433.hp1 HG01884.hp2 HG02109.hp2 others(19): Show |
synonymous_variant | LOW | c.204G>A | p.Gln68Gln | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 349/770 | 204/351 | 68/116 | chr1 | 152884626 |
| chr:pos | ref | alt | # # of thapid:transcript level |
thapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:152878430
|
G | A | 2 | a0001c0001t0002a0001c0001t0004 | 122 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(119): Show |
5_prime_UTR_premature_start_codon_gain_variant | LOW | c.-37G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/2 | chr1 | 152878430 | ||||||
| chr1:152884407
|
T | C | 1 | a0001c0001t0006 | 1 | HG02015.hp2 | 5_prime_UTR_variant | MODIFIER | c.-16T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 16 | chr1 | 152884407 | |||||
| chr1:152884412
|
G | C | 1 | a0001c0001t0007 | 1 | HG03486.hp1 | 5_prime_UTR_premature_start_codon_gain_variant | LOW | c.-11G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | chr1 | 152884412 | ||||||
| chr1:152884816
|
C | T | 2 | a0002c0002t0005a0002c0004t0005 | 2 | HG02809.hp2 HG03516.hp1 |
3_prime_UTR_variant | MODIFIER | c.*43C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 43 | chr1 | 152884816 | |||||
| chr1:152885005
|
C | A | 4 | a0001c0001t0004a0002c0002t0003a0002c0002t0005others(1): Show | 29 | HG01433.hp1 HG01884.hp2 HG02109.hp2 others(26): Show |
3_prime_UTR_variant | MODIFIER | c.*232C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 2/2 | 232 | chr1 | 152885005 |
| chr:pos | ref | alt | # # of ghapid:genebody level |
ghapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | genebody_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:152878451
|
C | A | 1 | a0001c0001t0001g0024 | 1 | HG02698.hp1 | splice_region_variant&intron_variant | LOW | c.-21+5C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878451 | ||||||
| chr1:152878462
|
A | C | 45 | a0001c0001t0001g0003a0001c0001t0001g0006a0001c0001t0001g0007others(42): Show | 220 | HG00280.hp2 HG00408.hp2 HG00438.hp1 others(217): Show |
intron_variant | MODIFIER | c.-21+16A>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878462 | ||||||
| chr1:152878542
|
A | G | 1 | a0001c0001t0001g0039 | 1 | HG02735.hp2 | intron_variant | MODIFIER | c.-21+96A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878542 | ||||||
| chr1:152878553
|
T | C | 1 | a0001c0001t0001g0025 | 1 | HG00609.hp1 | intron_variant | MODIFIER | c.-21+107T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878553 | ||||||
| chr1:152878652
|
A | G | 1 | a0002c0004t0005g0062 | 1 | HG03516.hp1 | intron_variant | MODIFIER | c.-21+206A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878652 | ||||||
| chr1:152878750
|
T | A | 1 | a0001c0001t0001g0040 | 1 | HG02965.hp1 | intron_variant | MODIFIER | c.-21+304T>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878750 | ||||||
| chr1:152878839
|
C | T | 15 | a0001c0001t0002g0002a0001c0001t0002g0004a0001c0001t0002g0016others(12): Show | 122 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(119): Show |
intron_variant | MODIFIER | c.-21+393C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878839 | ||||||
| chr1:152878939
|
G | T | 3 | a0001c0001t0001g0006a0001c0001t0001g0051a0001c0001t0006g0006 | 12 | HG00280.hp2 HG00733.hp1 HG00741.hp1 others(9): Show |
intron_variant | MODIFIER | c.-21+493G>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152878939 | ||||||
| chr1:152879111
|
C | T | 1 | a0001c0001t0001g0038 | 1 | HG00438.hp2 | intron_variant | MODIFIER | c.-21+665C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879111 | ||||||
| chr1:152879125
|
C | T | 1 | a0002c0002t0003g0050 | 1 | HG02922.hp2 | intron_variant | MODIFIER | c.-21+679C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879125 | ||||||
| chr1:152879222
|
C | A | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-21+776C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879222 | ||||||
| chr1:152879238
|
T | G | 1 | a0001c0001t0001g0040 | 1 | HG02965.hp1 | intron_variant | MODIFIER | c.-21+792T>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879238 | ||||||
| chr1:152879290
|
G | A | 1 | a0001c0001t0001g0044 | 1 | HG02145.hp1 | intron_variant | MODIFIER | c.-21+844G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879290 | ||||||
| chr1:152879331
|
C | T | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-21+885C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879331 | ||||||
| chr1:152879429
|
C | A | 1 | a0002c0002t0003g0041 | 1 | HG03579.hp2 | intron_variant | MODIFIER | c.-21+983C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879429 | ||||||
| chr1:152879798
|
G | T | 1 | a0001c0001t0001g0037 | 1 | HG00735.hp1 | intron_variant | MODIFIER | c.-21+1352G>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152879798 | ||||||
| chr1:152880039
|
C | A | 2 | a0001c0001t0002g0052a0001c0001t0002g0053 | 2 | HG00408.hp2 NA19012.hp2 |
intron_variant | MODIFIER | c.-21+1593C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880039 | ||||||
| chr1:152880364
|
G | A | 1 | a0002c0002t0003g0042 | 1 | HG02109.hp2 | intron_variant | MODIFIER | c.-21+1918G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880364 | ||||||
| chr1:152880540
|
G | A | 1 | a0001c0001t0001g0026 | 1 | HG01243.hp1 | intron_variant | MODIFIER | c.-21+2094G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880540 | ||||||
| chr1:152880633
|
C | T | 1 | a0001c0001t0001g0044 | 1 | HG02145.hp1 | intron_variant | MODIFIER | c.-21+2187C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880633 | ||||||
| chr1:152880661
|
T | C | 32 | a0001c0001t0001g0003a0001c0001t0001g0011a0001c0001t0001g0012others(29): Show | 175 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(172): Show |
intron_variant | MODIFIER | c.-21+2215T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880661 | ||||||
| chr1:152880715
|
T | A | 9 | a0001c0001t0001g0011a0002c0002t0003g0005a0002c0002t0003g0011others(6): Show | 23 | HG01433.hp1 HG02109.hp2 HG02280.hp1 others(20): Show |
intron_variant | MODIFIER | c.-21+2269T>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880715 | ||||||
| chr1:152880768
|
A | G | 1 | a0001c0001t0002g0061 | 1 | NA18975.hp1 | intron_variant | MODIFIER | c.-21+2322A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880768 | ||||||
| chr1:152880802
|
T | C | 14 | a0001c0001t0002g0002a0001c0001t0002g0004a0001c0001t0002g0016others(11): Show | 115 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(112): Show |
intron_variant | MODIFIER | c.-21+2356T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880802 | ||||||
| chr1:152880926
|
G | C | 32 | a0001c0001t0001g0003a0001c0001t0001g0011a0001c0001t0001g0012others(29): Show | 175 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(172): Show |
intron_variant | MODIFIER | c.-21+2480G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152880926 | ||||||
| chr1:152880979
|
C | CCCTCCTT others(79): Show |
1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-21+2542_-21+2543i others(88): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152880979 | |||||
| chr1:152880979
|
C | CCCTCCTT others(80): Show |
65 | a0001c0001t0001g0001a0001c0001t0001g0003a0001c0001t0001g0006others(62): Show | 416 | HG00140.hp1 HG00140.hp2 HG00280.hp1 others(413): Show |
intron_variant | MODIFIER | c.-21+2544_-21+2545i others(89): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152880979 | |||||
| chr1:152881068
|
G | A | 1 | a0001c0001t0002g0055 | 1 | HG04184.hp1 | intron_variant | MODIFIER | c.-21+2622G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881068 | ||||||
| chr1:152881078
|
G | A | 1 | a0001c0001t0001g0028 | 1 | HG02818.hp2 | intron_variant | MODIFIER | c.-21+2632G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881078 | ||||||
| chr1:152881110
|
G | A | 15 | a0001c0001t0002g0002a0001c0001t0002g0004a0001c0001t0002g0016others(12): Show | 122 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(119): Show |
intron_variant | MODIFIER | c.-21+2664G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881110 | ||||||
| chr1:152881130
|
G | C | 1 | a0001c0001t0001g0018 | 2 | HG01243.hp2 HG01934.hp2 |
intron_variant | MODIFIER | c.-21+2684G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881130 | ||||||
| chr1:152881268
|
C | A | 1 | a0001c0001t0001g0040 | 1 | HG02965.hp1 | intron_variant | MODIFIER | c.-21+2822C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881268 | ||||||
| chr1:152881377
|
C | A | 1 | a0001c0001t0001g0014 | 4 | HG02895.hp1 HG03139.hp1 HG03209.hp1 others(1): Show |
intron_variant | MODIFIER | c.-21+2931C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881377 | ||||||
| chr1:152881410
|
C | T | 1 | a0001c0001t0001g0044 | 1 | HG02145.hp1 | intron_variant | MODIFIER | c.-21+2964C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881410 | ||||||
| chr1:152881427
|
G | T | 1 | a0001c0001t0001g0036 | 1 | NA18957.hp2 | intron_variant | MODIFIER | c.-20-2976G>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881427 | ||||||
| chr1:152881459
|
C | A | 1 | a0001c0001t0001g0029 | 1 | NA19090.hp1 | intron_variant | MODIFIER | c.-20-2944C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881459 | ||||||
| chr1:152881459
|
C | T | 9 | a0001c0001t0001g0011a0002c0002t0003g0005a0002c0002t0003g0011others(6): Show | 23 | HG01433.hp1 HG02109.hp2 HG02280.hp1 others(20): Show |
intron_variant | MODIFIER | c.-20-2944C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881459 | ||||||
| chr1:152881550
|
T | C | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-2853T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881550 | ||||||
| chr1:152881561
|
G | A | 1 | a0001c0001t0001g0020 | 2 | HG01975.hp1 HG02647.hp1 |
intron_variant | MODIFIER | c.-20-2842G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881561 | ||||||
| chr1:152881565
|
G | C | 15 | a0001c0001t0002g0002a0001c0001t0002g0004a0001c0001t0002g0016others(12): Show | 122 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(119): Show |
intron_variant | MODIFIER | c.-20-2838G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881565 | ||||||
| chr1:152881637
|
G | A | 9 | a0001c0001t0001g0011a0002c0002t0003g0005a0002c0002t0003g0011others(6): Show | 23 | HG01433.hp1 HG02109.hp2 HG02280.hp1 others(20): Show |
intron_variant | MODIFIER | c.-20-2766G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881637 | ||||||
| chr1:152881646
|
C | T | 1 | a0001c0001t0001g0035 | 1 | NA19075.hp2 | intron_variant | MODIFIER | c.-20-2757C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881646 | ||||||
| chr1:152881678
|
G | A | 1 | a0003c0003t0001g0047 | 1 | HG02630.hp1 | intron_variant | MODIFIER | c.-20-2725G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881678 | ||||||
| chr1:152881681
|
C | T | 1 | a0001c0001t0001g0046 | 1 | homoSapiens_chm13v2.hp1 | intron_variant | MODIFIER | c.-20-2722C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881681 | ||||||
| chr1:152881694
|
C | CA | 5 | a0001c0001t0001g0008a0001c0001t0001g0015a0001c0001t0001g0017others(2): Show | 17 | HG02451.hp2 HG02486.hp1 HG02647.hp2 others(14): Show |
intron_variant | MODIFIER | c.-20-2690dupA | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881694 | |||||
| chr1:152881694
|
C | CAA | 8 | a0001c0001t0001g0012a0001c0001t0001g0013a0001c0001t0001g0021others(5): Show | 16 | HG01884.hp2 HG01891.hp1 HG02055.hp1 others(13): Show |
intron_variant | MODIFIER | c.-20-2691_-20-2690d others(4): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881694 | |||||
| chr1:152881694
|
C | CAAA | 13 | a0001c0001t0001g0003a0001c0001t0001g0011a0001c0001t0001g0020others(10): Show | 37 | HG00558.hp2 HG00609.hp2 HG01069.hp2 others(34): Show |
intron_variant | MODIFIER | c.-20-2692_-20-2690d others(5): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881694 | |||||
| chr1:152881694
|
C | CAAAA | 14 | a0001c0001t0002g0002a0001c0001t0002g0023a0001c0001t0002g0052others(11): Show | 117 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(114): Show |
intron_variant | MODIFIER | c.-20-2693_-20-2690d others(6): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881694 | |||||
| chr1:152881694
|
C | CAAAAA | 2 | a0001c0001t0002g0004a0001c0001t0002g0054 | 14 | HG01106.hp1 HG02056.hp1 HG02738.hp1 others(11): Show |
intron_variant | MODIFIER | c.-20-2694_-20-2690d others(7): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881694 | |||||
| chr1:152881717
|
A | G | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-2686A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881717 | ||||||
| chr1:152881784
|
A | G | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-2619A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881784 | ||||||
| chr1:152881825
|
AGGACCTT others(11): Show |
A | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2575_-20-2558d others(20): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881825 | |||||
| chr1:152881865
|
C | G | 1 | a0001c0001t0001g0039 | 1 | HG02735.hp2 | intron_variant | MODIFIER | c.-20-2538C>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881865 | ||||||
| chr1:152881910
|
T | G | 2 | a0001c0001t0001g0021a0001c0001t0001g0044 | 3 | HG01891.hp1 HG02109.hp1 HG02145.hp1 |
intron_variant | MODIFIER | c.-20-2493T>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881910 | ||||||
| chr1:152881926
|
A | T | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2477A>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881926 | ||||||
| chr1:152881938
|
G | T | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2465G>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881938 | ||||||
| chr1:152881952
|
A | G | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2451A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881952 | ||||||
| chr1:152881953
|
A | ATTATT | 12 | a0001c0001t0002g0002a0001c0001t0002g0004a0001c0001t0002g0016others(9): Show | 113 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(110): Show |
intron_variant | MODIFIER | c.-20-2431_-20-2427d others(7): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881953 | |||||
| chr1:152881953
|
A | ATTATTTT others(3): Show |
1 | a0001c0001t0002g0053 | 1 | NA19012.hp2 | intron_variant | MODIFIER | c.-20-2436_-20-2427d others(12): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152881953 | |||||
| chr1:152881957
|
T | C | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2446T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881957 | ||||||
| chr1:152881961
|
A | G | 9 | a0001c0001t0001g0011a0002c0002t0003g0005a0002c0002t0003g0011others(6): Show | 23 | HG01433.hp1 HG02109.hp2 HG02280.hp1 others(20): Show |
intron_variant | MODIFIER | c.-20-2442A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881961 | ||||||
| chr1:152881965
|
T | A | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2438T>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881965 | ||||||
| chr1:152881970
|
T | TCAGTCTT others(5): Show |
1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2433_-20-2432i others(14): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881970 | ||||||
| chr1:152881971
|
A | G | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2432A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881971 | ||||||
| chr1:152881981
|
A | T | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2422A>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881981 | ||||||
| chr1:152881986
|
A | T | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2417A>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881986 | ||||||
| chr1:152881997
|
T | C | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2406T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152881997 | ||||||
| chr1:152882011
|
T | G | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2392T>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882011 | ||||||
| chr1:152882012
|
C | T | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2391C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882012 | ||||||
| chr1:152882013
|
T | C | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2390T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882013 | ||||||
| chr1:152882019
|
A | C | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2384A>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882019 | ||||||
| chr1:152882027
|
G | C | 1 | a0001c0001t0002g0054 | 1 | NA19001.hp1 | intron_variant | MODIFIER | c.-20-2376G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882027 | ||||||
| chr1:152882118
|
G | A | 1 | a0001c0001t0001g0049 | 1 | NA19043.hp2 | intron_variant | MODIFIER | c.-20-2285G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882118 | ||||||
| chr1:152882124
|
T | C | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-2279T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882124 | ||||||
| chr1:152882177
|
G | C | 1 | a0001c0001t0001g0040 | 1 | HG02965.hp1 | intron_variant | MODIFIER | c.-20-2226G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882177 | ||||||
| chr1:152882195
|
C | T | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-2208C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882195 | ||||||
| chr1:152882251
|
G | A | 1 | a0001c0001t0001g0019 | 2 | HG00558.hp1 NA19078.hp2 |
intron_variant | MODIFIER | c.-20-2152G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882251 | ||||||
| chr1:152882266
|
C | T | 2 | a0002c0002t0003g0005a0002c0002t0003g0042 | 13 | HG01433.hp1 HG02109.hp2 HG02280.hp2 others(10): Show |
intron_variant | MODIFIER | c.-20-2137C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882266 | ||||||
| chr1:152882288
|
T | G | 3 | a0001c0001t0001g0012a0001c0001t0001g0045a0001c0001t0007g0012 | 6 | HG02055.hp1 HG02258.hp2 HG02622.hp1 others(3): Show |
intron_variant | MODIFIER | c.-20-2115T>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882288 | ||||||
| chr1:152882448
|
A | G | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-1955A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882448 | ||||||
| chr1:152882462
|
G | A | 1 | a0002c0002t0003g0050 | 1 | HG02922.hp2 | intron_variant | MODIFIER | c.-20-1941G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882462 | ||||||
| chr1:152882557
|
C | T | 2 | a0001c0001t0001g0012a0001c0001t0007g0012 | 5 | HG02055.hp1 HG02258.hp2 HG02895.hp2 others(2): Show |
intron_variant | MODIFIER | c.-20-1846C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882557 | ||||||
| chr1:152882631
|
A | G | 1 | a0001c0001t0002g0060 | 1 | NA19077.hp2 | intron_variant | MODIFIER | c.-20-1772A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882631 | ||||||
| chr1:152882656
|
G | C | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-1747G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882656 | ||||||
| chr1:152882659
|
G | GCA | 11 | a0001c0001t0001g0003a0001c0001t0001g0010a0001c0001t0001g0011others(8): Show | 40 | HG00609.hp2 HG01975.hp1 HG02055.hp1 others(37): Show |
intron_variant | MODIFIER | c.-20-1722_-20-1721d others(4): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152882659 | |||||
| chr1:152882659
|
G | GCACA | 5 | a0001c0001t0001g0021a0001c0001t0001g0044a0002c0002t0003g0005others(2): Show | 17 | HG01433.hp1 HG01891.hp1 HG02109.hp1 others(14): Show |
intron_variant | MODIFIER | c.-20-1724_-20-1721d others(6): Show |
SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | INFO_REALIGN_3_PRIME | chr1 | 152882659 | |||||
| chr1:152882677
|
A | G | 1 | a0001c0001t0002g0059 | 1 | NA19062.hp1 | intron_variant | MODIFIER | c.-20-1726A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882677 | ||||||
| chr1:152882796
|
C | A | 1 | a0001c0001t0002g0056 | 1 | NA18975.hp2 | intron_variant | MODIFIER | c.-20-1607C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882796 | ||||||
| chr1:152882835
|
G | A | 1 | a0001c0001t0001g0015 | 4 | HG02717.hp1 HG02818.hp1 NA19240.hp1 others(1): Show |
intron_variant | MODIFIER | c.-20-1568G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882835 | ||||||
| chr1:152882957
|
T | G | 1 | a0001c0001t0001g0048 | 1 | HG03486.hp2 | intron_variant | MODIFIER | c.-20-1446T>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882957 | ||||||
| chr1:152882967
|
C | G | 1 | a0001c0001t0001g0034 | 1 | NA18982.hp1 | intron_variant | MODIFIER | c.-20-1436C>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152882967 | ||||||
| chr1:152883160
|
C | G | 2 | a0001c0001t0002g0058a0001c0001t0002g0060 | 2 | NA18959.hp2 NA19077.hp2 |
intron_variant | MODIFIER | c.-20-1243C>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883160 | ||||||
| chr1:152883175
|
A | C | 1 | a0002c0004t0005g0062 | 1 | HG03516.hp1 | intron_variant | MODIFIER | c.-20-1228A>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883175 | ||||||
| chr1:152883228
|
G | A | 2 | a0001c0001t0001g0021a0001c0001t0001g0044 | 3 | HG01891.hp1 HG02109.hp1 HG02145.hp1 |
intron_variant | MODIFIER | c.-20-1175G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883228 | ||||||
| chr1:152883270
|
C | T | 1 | a0002c0002t0003g0041 | 1 | HG03579.hp2 | intron_variant | MODIFIER | c.-20-1133C>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883270 | ||||||
| chr1:152883386
|
G | A | 2 | a0002c0002t0003g0005a0002c0002t0003g0042 | 13 | HG01433.hp1 HG02109.hp2 HG02280.hp2 others(10): Show |
intron_variant | MODIFIER | c.-20-1017G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883386 | ||||||
| chr1:152883421
|
G | A | 1 | a0001c0001t0002g0023 | 2 | NA18945.hp1 NA19080.hp1 |
intron_variant | MODIFIER | c.-20-982G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883421 | ||||||
| chr1:152883425
|
G | A | 1 | a0001c0001t0001g0036 | 1 | NA18957.hp2 | intron_variant | MODIFIER | c.-20-978G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883425 | ||||||
| chr1:152883476
|
G | A | 1 | a0001c0001t0001g0037 | 1 | HG00735.hp1 | intron_variant | MODIFIER | c.-20-927G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883476 | ||||||
| chr1:152883479
|
C | A | 1 | a0001c0001t0001g0030 | 1 | NA19068.hp1 | intron_variant | MODIFIER | c.-20-924C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883479 | ||||||
| chr1:152883564
|
C | G | 1 | a0001c0001t0001g0051 | 1 | HG00280.hp2 | intron_variant | MODIFIER | c.-20-839C>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883564 | ||||||
| chr1:152883608
|
G | C | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-795G>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883608 | ||||||
| chr1:152883611
|
G | T | 1 | a0001c0001t0001g0033 | 1 | NA19010.hp2 | intron_variant | MODIFIER | c.-20-792G>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883611 | ||||||
| chr1:152883697
|
G | A | 2 | a0001c0001t0001g0012a0001c0001t0007g0012 | 5 | HG02055.hp1 HG02258.hp2 HG02895.hp2 others(2): Show |
intron_variant | MODIFIER | c.-20-706G>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883697 | ||||||
| chr1:152883740
|
T | A | 1 | a0001c0001t0001g0028 | 1 | HG02818.hp2 | intron_variant | MODIFIER | c.-20-663T>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883740 | ||||||
| chr1:152883826
|
A | G | 24 | a0001c0001t0001g0011a0001c0001t0002g0002a0001c0001t0002g0004others(21): Show | 145 | HG00408.hp2 HG00438.hp1 HG00544.hp1 others(142): Show |
intron_variant | MODIFIER | c.-20-577A>G | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883826 | ||||||
| chr1:152883864
|
T | C | 1 | a0001c0001t0002g0057 | 1 | NA18985.hp1 | intron_variant | MODIFIER | c.-20-539T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883864 | ||||||
| chr1:152883885
|
A | C | 1 | a0001c0001t0001g0022 | 2 | HG02965.hp2 NA20300.hp1 |
intron_variant | MODIFIER | c.-20-518A>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883885 | ||||||
| chr1:152883930
|
C | A | 8 | a0001c0001t0001g0011a0002c0002t0003g0005a0002c0002t0003g0011others(5): Show | 22 | HG01433.hp1 HG02109.hp2 HG02280.hp1 others(19): Show |
intron_variant | MODIFIER | c.-20-473C>A | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883930 | ||||||
| chr1:152883999
|
G | T | 1 | a0001c0001t0001g0032 | 1 | HG03239.hp1 | intron_variant | MODIFIER | c.-20-404G>T | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152883999 | ||||||
| chr1:152884031
|
T | C | 1 | a0001c0001t0001g0031 | 1 | NA18943.hp1 | intron_variant | MODIFIER | c.-20-372T>C | SMCP | ENSG00000163206.6 | transcript | ENST00000368765.4 | protein_coding | 1/1 | chr1 | 152884031 |
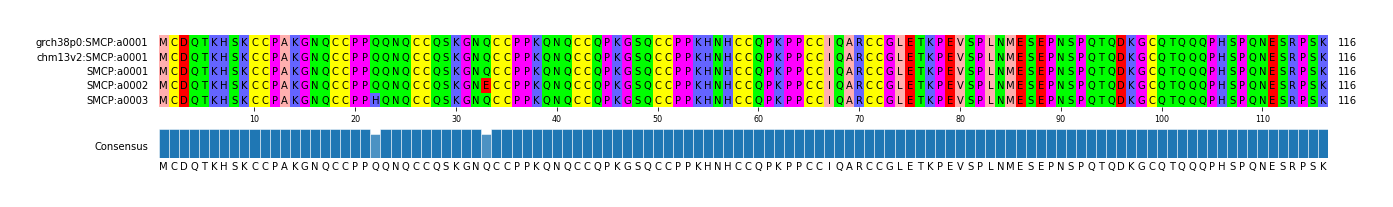 Multiple Alignments (by aa haplotype frequency)
Multiple Alignments (by aa haplotype frequency)
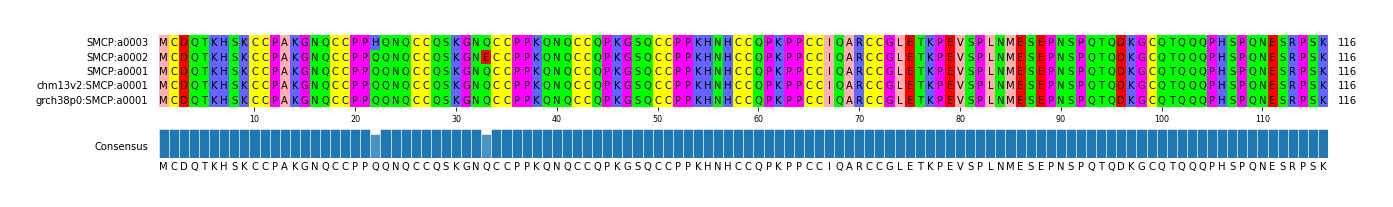 Multiple Alignments (by aa haplotype similarity)
Multiple Alignments (by aa haplotype similarity)