Search by Genic region
| geneid | 93190 |
|---|---|
| ensemblid | ENSG00000157330.10 |
| hgncid | 28567 |
| symbol | CFAP107 |
| name | cilia and flagella associated protein 107 |
| refseq_nuc | NM_152290.4 |
| refseq_prot | NP_689503.3 |
| ensembl_nuc | ENST00000614859.5 |
| ensembl_prot | ENSP00000477802.1 |
| mane_status | MANE Select |
| chr | chr1 |
| start | 12746200 |
| end | 12763699 |
| strand | + |
| ver | v1.2 |
| region | chr1:12746200-12763699 |
| region5000 | chr1:12741200-12768699 |
| regionname0 | CFAP107_chr1_12746200_12763699 |
| regionname5000 | CFAP107_chr1_12741200_12768699 |
| ahapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
alen | total | AFR | AMR | EAS | EUR | SAS | JPT | regionname | genename | aa | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001 | 1/0 | 194 | 249 | 67 | 43 | 94 | 13 | 31 | 75 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002 | 0/1 | 194 | 68 | 7 | 15 | 32 | 4 | 9 | 25 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0003 | 0/0 | 194 | 5 | 4 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0004 | 0/0 | 194 | 5 | 5 | 0 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0005 | 0/0 | 194 | 5 | 5 | 0 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0006 | 0/0 | 194 | 2 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0007 | 0/0 | 33 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0008 | 0/0 | 13 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| chapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
clen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | cseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| c0001 | 1/0 | 585 | 247 | 66 | 43 | 94 | 13 | 30 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0002 | 0/1 | 585 | 68 | 7 | 15 | 32 | 4 | 9 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0003 | 0/0 | 585 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0004 | 0/0 | 585 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0005 | 0/0 | 585 | 5 | 4 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0006 | 0/0 | 585 | 2 | 1 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0007 | 0/0 | 585 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0008 | 0/0 | 585 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0009 | 0/0 | 585 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| c0010 | 0/0 | 474 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| thapid | grch38chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
tlen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | tseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| t0001 | 0/0 | 2981 | 85 | 32 | 19 | 23 | 2 | 9 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0002 | 1/0 | 2980 | 69 | 1 | 18 | 35 | 3 | 11 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0003 | 0/0 | 3195 | 59 | 6 | 13 | 31 | 3 | 6 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0004 | 0/0 | 2981 | 14 | 9 | 1 | 1 | 0 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0005 | 0/0 | 2982 | 13 | 0 | 0 | 13 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0006 | 0/0 | 2981 | 13 | 2 | 1 | 8 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0007 | 0/0 | 2982 | 9 | 0 | 0 | 9 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0008 | 0/0 | 2981 | 7 | 6 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0009 | 0/1 | 3195 | 6 | 0 | 2 | 0 | 1 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0010 | 0/0 | 2980 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0011 | 0/0 | 2982 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0012 | 0/0 | 2982 | 4 | 3 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0013 | 0/0 | 2982 | 4 | 1 | 0 | 0 | 2 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0014 | 0/0 | 2981 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0015 | 0/0 | 2981 | 3 | 0 | 1 | 0 | 2 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0016 | 0/0 | 2981 | 2 | 0 | 2 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0017 | 0/0 | 2981 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0018 | 0/0 | 2981 | 2 | 1 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0019 | 0/0 | 2980 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0020 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0021 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0022 | 0/0 | 3407 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0023 | 0/0 | 3407 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0024 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0025 | 0/0 | 2981 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0026 | 0/0 | 3194 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0027 | 0/0 | 2981 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0028 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0029 | 0/0 | 2980 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0030 | 0/0 | 2982 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0031 | 0/0 | 2981 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0032 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0033 | 0/0 | 2981 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0034 | 0/0 | 2981 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0035 | 0/0 | 3193 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0036 | 0/0 | 3193 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0037 | 0/0 | 2980 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0038 | 0/0 | 2980 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0039 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0040 | 0/0 | 2980 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0041 | 0/0 | 2982 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0042 | 0/0 | 3195 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0043 | 0/0 | 3407 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0044 | 0/0 | 3193 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0045 | 0/0 | 2981 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0046 | 0/0 | 2981 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0047 | 0/0 | 2982 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0048 | 0/0 | 2982 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| t0049 | 0/0 | 2749 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| ghapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
total | AFR | AMR | EAS | EUR | SAS | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| g0001 | 1/0 | 52 | 2 | 9 | 29 | 7 | 4 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0002 | 0/0 | 44 | 10 | 6 | 17 | 3 | 8 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0003 | 0/0 | 39 | 2 | 8 | 22 | 1 | 6 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0004 | 0/0 | 20 | 1 | 0 | 17 | 1 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0005 | 0/0 | 15 | 8 | 2 | 1 | 1 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0006 | 0/0 | 7 | 2 | 1 | 3 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0007 | 0/0 | 5 | 4 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0008 | 0/0 | 4 | 0 | 3 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0009 | 0/0 | 4 | 1 | 3 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0010 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0011 | 0/0 | 3 | 2 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0012 | 0/0 | 3 | 0 | 3 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0013 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0014 | 0/0 | 3 | 0 | 3 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0015 | 0/0 | 3 | 0 | 0 | 0 | 0 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0016 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0017 | 0/0 | 3 | 1 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0018 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0019 | 0/0 | 3 | 2 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0020 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0021 | 0/0 | 2 | 0 | 2 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0022 | 0/0 | 2 | 0 | 2 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0023 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0024 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0025 | 0/0 | 2 | 0 | 1 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0026 | 0/0 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0027 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0028 | 0/0 | 2 | 0 | 0 | 1 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0029 | 0/0 | 2 | 0 | 1 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0030 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0031 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0032 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0033 | 0/0 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0034 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0035 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0036 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0037 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0038 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0039 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0040 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0041 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0042 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0043 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0044 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0045 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0046 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0047 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0048 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0049 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0050 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0051 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0052 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0053 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0054 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0055 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0056 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0057 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0058 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0059 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0060 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0061 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0062 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0063 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0064 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0065 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0066 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0067 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0068 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0069 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0070 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0071 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0072 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0073 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0074 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0075 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0076 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0077 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0078 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0079 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0080 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0081 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0082 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0083 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0084 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0085 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0086 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0087 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0088 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0089 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0090 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0091 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0092 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0093 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0094 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0095 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0096 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0097 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0098 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0099 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0100 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0101 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0102 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0103 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0104 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0105 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0106 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0107 | 0/1 | 1 | 0 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0108 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0109 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0110 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0111 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0112 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0113 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0114 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0115 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0116 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0117 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0118 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0119 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| g0120 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| achapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
clen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | cseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001 | 1/0 | 585 | 247 | 66 | 43 | 94 | 13 | 30 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0007 | 0/0 | 585 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0008 | 0/0 | 585 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002c0002 | 0/1 | 585 | 68 | 7 | 15 | 32 | 4 | 9 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0003c0005 | 0/0 | 585 | 5 | 4 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0004c0004 | 0/0 | 585 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0005c0003 | 0/0 | 585 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0006c0006 | 0/0 | 585 | 2 | 1 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0007c0009 | 0/0 | 585 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0008c0010 | 0/0 | 474 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| acthapid | grch38chm13v2 | tlen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | tseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001t0001 | 0/0 | 3565 | 83 | 31 | 19 | 23 | 2 | 8 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0002 | 1/0 | 3564 | 68 | 1 | 17 | 35 | 3 | 11 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0004 | 0/0 | 3565 | 14 | 9 | 1 | 1 | 0 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0005 | 0/0 | 3566 | 13 | 0 | 0 | 13 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0006 | 0/0 | 3565 | 13 | 2 | 1 | 8 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0007 | 0/0 | 3566 | 9 | 0 | 0 | 9 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0008 | 0/0 | 3565 | 7 | 6 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0012 | 0/0 | 3566 | 4 | 3 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0013 | 0/0 | 3566 | 4 | 1 | 0 | 0 | 2 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0015 | 0/0 | 3565 | 3 | 0 | 1 | 0 | 2 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0016 | 0/0 | 3565 | 2 | 0 | 2 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0017 | 0/0 | 3565 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0019 | 0/0 | 3564 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0020 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0021 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0024 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0025 | 0/0 | 3565 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0026 | 0/0 | 3778 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0027 | 0/0 | 3565 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0028 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0029 | 0/0 | 3564 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0030 | 0/0 | 3566 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0031 | 0/0 | 3565 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0032 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0033 | 0/0 | 3565 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0034 | 0/0 | 3565 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0036 | 0/0 | 3777 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0037 | 0/0 | 3564 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0038 | 0/0 | 3564 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0039 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0040 | 0/0 | 3564 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0041 | 0/0 | 3566 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0044 | 0/0 | 3777 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0045 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0046 | 0/0 | 3565 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0047 | 0/0 | 3566 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0001t0048 | 0/0 | 3566 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0007t0001 | 0/0 | 3565 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0001c0008t0001 | 0/0 | 3565 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002c0002t0003 | 0/0 | 3779 | 59 | 6 | 13 | 31 | 3 | 6 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002c0002t0009 | 0/1 | 3779 | 6 | 0 | 2 | 0 | 1 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002c0002t0035 | 0/0 | 3777 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002c0002t0042 | 0/0 | 3779 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0002c0002t0043 | 0/0 | 3991 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0003c0005t0014 | 0/0 | 3565 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0003c0005t0022 | 0/0 | 3991 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0003c0005t0023 | 0/0 | 3991 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0004c0004t0010 | 0/0 | 3564 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0005c0003t0011 | 0/0 | 3566 | 5 | 5 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0006c0006t0018 | 0/0 | 3565 | 2 | 1 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0007c0009t0002 | 0/0 | 3564 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| a0008c0010t0049 | 0/0 | 3222 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | copy fasta | chr1 | 12741200 | 12768699 |
| actghapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
total | AFR | AMR | EAS | EUR | SAS | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001t0001g0001 | 0/0 | 9 | 0 | 3 | 4 | 2 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0002 | 0/0 | 26 | 9 | 5 | 9 | 0 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0006 | 0/0 | 5 | 2 | 0 | 2 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0007 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0009 | 0/0 | 4 | 1 | 3 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0011 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0013 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0014 | 0/0 | 3 | 0 | 3 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0015 | 0/0 | 3 | 0 | 0 | 0 | 0 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0016 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0023 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0038 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0049 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0050 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0052 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0053 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0055 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0058 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0059 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0061 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0062 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0063 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0065 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0069 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0071 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0072 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0073 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0074 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0076 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0080 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0085 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0088 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0001g0112 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0001 | 1/0 | 34 | 1 | 6 | 20 | 2 | 4 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0002 | 0/0 | 6 | 0 | 0 | 5 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0012 | 0/0 | 3 | 0 | 3 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0021 | 0/0 | 2 | 0 | 2 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0022 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0024 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0025 | 0/0 | 2 | 0 | 1 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0033 | 0/0 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0036 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0037 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0041 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0042 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0048 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0054 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0057 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0067 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0075 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0077 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0078 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0079 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0083 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0084 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0086 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0002g0087 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0004g0005 | 0/0 | 12 | 7 | 1 | 1 | 0 | 3 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0004g0040 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0004g0046 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0005g0004 | 0/0 | 9 | 0 | 0 | 9 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0005g0027 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0005g0093 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0005g0094 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0001 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0002 | 0/0 | 4 | 1 | 1 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0006 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0011 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0026 | 0/0 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0047 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0056 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0006g0082 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0007g0004 | 0/0 | 7 | 0 | 0 | 7 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0007g0028 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0007g0091 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0008g0007 | 0/0 | 4 | 3 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0008g0060 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0008g0064 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0008g0081 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0012g0019 | 0/0 | 3 | 2 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0012g0115 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0013g0004 | 0/0 | 2 | 1 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0013g0028 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0013g0092 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0015g0002 | 0/0 | 2 | 0 | 0 | 0 | 2 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0015g0006 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0016g0005 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0016g0039 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0017g0002 | 0/0 | 2 | 0 | 0 | 0 | 0 | 2 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0019g0117 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0020g0114 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0021g0116 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0024g0113 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0025g0004 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0026g0108 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0027g0005 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0028g0005 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0029g0045 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0030g0004 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0031g0002 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0032g0066 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0033g0002 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0034g0001 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0036g0001 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0037g0001 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0038g0001 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0039g0051 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0040g0002 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0041g0001 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0044g0109 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0045g0096 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0046g0002 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0047g0110 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0001t0048g0111 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0007t0001g0089 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0001c0008t0001g0070 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0003 | 0/0 | 34 | 2 | 6 | 21 | 0 | 5 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0008 | 0/0 | 4 | 0 | 3 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0017 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0029 | 0/0 | 2 | 0 | 1 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0030 | 0/0 | 2 | 0 | 0 | 2 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0031 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0043 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0044 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0068 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0090 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0097 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0098 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0099 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0100 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0101 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0102 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0103 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0104 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0003g0105 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0009g0003 | 0/0 | 4 | 0 | 2 | 0 | 1 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0009g0106 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0009g0107 | 0/1 | 1 | 0 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0035g0095 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0042g0017 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0002c0002t0043g0003 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0003c0005t0014g0010 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0003c0005t0022g0034 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0003c0005t0023g0035 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0004c0004t0010g0020 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0004c0004t0010g0118 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0004c0004t0010g0119 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0005c0003t0011g0018 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0005c0003t0011g0032 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0006c0006t0018g0001 | 0/0 | 2 | 1 | 0 | 0 | 1 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0007c0009t0002g0022 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| a0008c0010t0049g0120 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| sampleid | ID haplotypeid
|
ahapid | chapid | thapid | ghapid | gpopname | popname | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| HG00099 | hp1 | a0001 | c0001 | t0002 | g0084 | EUR | GBR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00099 | hp2 | a0001 | c0001 | t0002 | g0001 | EUR | GBR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00140 | hp1 | a0006 | c0006 | t0018 | g0001 | EUR | GBR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00140 | hp2 | a0001 | c0001 | t0031 | g0002 | EUR | GBR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00280 | hp1 | a0001 | c0001 | t0041 | g0001 | EUR | FIN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00280 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | FIN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00323 | hp1 | a0002 | c0002 | t0003 | g0097 | EUR | FIN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00323 | hp2 | a0002 | c0002 | t0003 | g0044 | EUR | FIN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00423 | hp1 | a0002 | c0002 | t0003 | g0103 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00423 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00438 | hp1 | a0002 | c0002 | t0003 | g0030 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00438 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00544 | hp1 | a0002 | c0002 | t0043 | g0003 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00544 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00597 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00597 | hp2 | a0001 | c0001 | t0005 | g0004 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00609 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00609 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | CHS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00639 | hp1 | a0002 | c0002 | t0003 | g0043 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00639 | hp2 | a0001 | c0001 | t0001 | g0009 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00642 | hp1 | a0003 | c0005 | t0022 | g0034 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00642 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00735 | hp1 | a0001 | c0001 | t0002 | g0012 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00735 | hp2 | a0001 | c0001 | t0001 | g0063 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00738 | hp1 | a0001 | c0001 | t0001 | g0009 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00738 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00741 | hp1 | a0001 | c0001 | t0001 | g0009 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG00741 | hp2 | a0001 | c0001 | t0001 | g0052 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01069 | hp1 | a0002 | c0002 | t0003 | g0008 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01069 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01070 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01070 | hp2 | a0002 | c0002 | t0009 | g0003 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01081 | hp1 | a0001 | c0001 | t0001 | g0080 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01081 | hp2 | a0001 | c0001 | t0002 | g0078 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01106 | hp1 | a0001 | c0001 | t0002 | g0001 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01106 | hp2 | a0001 | c0001 | t0016 | g0039 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01109 | hp1 | a0001 | c0001 | t0012 | g0019 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01109 | hp2 | a0002 | c0002 | t0003 | g0029 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01167 | hp1 | a0002 | c0002 | t0003 | g0003 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01167 | hp2 | a0001 | c0001 | t0008 | g0007 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01168 | hp1 | a0002 | c0002 | t0003 | g0008 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01168 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01169 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01169 | hp2 | a0002 | c0002 | t0003 | g0003 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01175 | hp1 | a0002 | c0002 | t0003 | g0003 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01175 | hp2 | a0002 | c0002 | t0003 | g0098 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01192 | hp1 | a0001 | c0001 | t0016 | g0005 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01192 | hp2 | a0001 | c0001 | t0006 | g0002 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01243 | hp1 | a0001 | c0001 | t0004 | g0005 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01243 | hp2 | a0001 | c0001 | t0002 | g0001 | AMR | PUR | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01255 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01255 | hp2 | a0002 | c0002 | t0009 | g0003 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01256 | hp1 | a0001 | c0001 | t0002 | g0012 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01256 | hp2 | a0001 | c0001 | t0001 | g0055 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01258 | hp1 | a0001 | c0001 | t0002 | g0012 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01258 | hp2 | a0001 | c0001 | t0002 | g0001 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01261 | hp1 | a0007 | c0009 | t0002 | g0022 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01261 | hp2 | a0001 | c0001 | t0002 | g0022 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01358 | hp1 | a0001 | c0001 | t0015 | g0006 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01358 | hp2 | a0002 | c0002 | t0003 | g0099 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01361 | hp1 | a0001 | c0001 | t0002 | g0042 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01361 | hp2 | a0002 | c0002 | t0003 | g0003 | AMR | CLM | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01515 | hp1 | a0001 | c0001 | t0015 | g0002 | EUR | IBS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01515 | hp2 | a0002 | c0002 | t0003 | g0008 | EUR | IBS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01516 | hp1 | a0001 | c0001 | t0013 | g0028 | EUR | IBS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01516 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | IBS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01517 | hp1 | a0001 | c0001 | t0013 | g0004 | EUR | IBS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01517 | hp2 | a0001 | c0001 | t0015 | g0002 | EUR | IBS | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01884 | hp1 | a0001 | c0001 | t0004 | g0005 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01884 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01928 | hp1 | a0001 | c0001 | t0001 | g0014 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01928 | hp2 | a0001 | c0001 | t0001 | g0074 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01934 | hp1 | a0001 | c0001 | t0001 | g0014 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01934 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01952 | hp1 | a0002 | c0002 | t0003 | g0008 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01952 | hp2 | a0001 | c0001 | t0002 | g0083 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01975 | hp1 | a0002 | c0002 | t0003 | g0003 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01975 | hp2 | a0001 | c0001 | t0002 | g0025 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01978 | hp1 | a0001 | c0001 | t0002 | g0021 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01978 | hp2 | a0002 | c0002 | t0003 | g0003 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01981 | hp1 | a0001 | c0001 | t0002 | g0075 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG01981 | hp2 | a0001 | c0001 | t0002 | g0001 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02055 | hp1 | a0001 | c0001 | t0001 | g0059 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02055 | hp2 | a0005 | c0003 | t0011 | g0018 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02071 | hp1 | a0001 | c0001 | t0005 | g0004 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02071 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02074 | hp1 | a0001 | c0001 | t0006 | g0006 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02074 | hp2 | a0001 | c0001 | t0002 | g0067 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02132 | hp1 | a0001 | c0001 | t0002 | g0024 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02132 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02145 | hp1 | a0001 | c0001 | t0004 | g0046 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02145 | hp2 | a0001 | c0001 | t0001 | g0006 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02148 | hp1 | a0001 | c0001 | t0002 | g0001 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02148 | hp2 | a0001 | c0001 | t0001 | g0014 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02155 | hp1 | a0001 | c0001 | t0006 | g0002 | EAS | CDX | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02155 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | CDX | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02165 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CDX | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02165 | hp2 | a0001 | c0001 | t0001 | g0038 | EAS | CDX | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02257 | hp1 | a0004 | c0004 | t0010 | g0118 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02257 | hp2 | a0001 | c0001 | t0001 | g0006 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02258 | hp1 | a0002 | c0002 | t0003 | g0068 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02258 | hp2 | a0001 | c0001 | t0001 | g0088 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02280 | hp1 | a0001 | c0001 | t0004 | g0005 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02280 | hp2 | a0001 | c0001 | t0012 | g0115 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02300 | hp1 | a0001 | c0001 | t0002 | g0021 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02300 | hp2 | a0001 | c0001 | t0002 | g0001 | AMR | PEL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02451 | hp1 | a0001 | c0001 | t0044 | g0109 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02451 | hp2 | a0001 | c0001 | t0013 | g0004 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02523 | hp1 | a0001 | c0001 | t0001 | g0023 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02523 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | KHV | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02572 | hp1 | a0001 | c0001 | t0004 | g0040 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02572 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02602 | hp1 | a0002 | c0002 | t0003 | g0003 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02602 | hp2 | a0001 | c0001 | t0001 | g0015 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02615 | hp1 | a0001 | c0001 | t0001 | g0016 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02615 | hp2 | a0001 | c0001 | t0001 | g0013 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02622 | hp1 | a0001 | c0001 | t0001 | g0013 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02622 | hp2 | a0001 | c0001 | t0008 | g0007 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02630 | hp1 | a0001 | c0001 | t0004 | g0005 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02630 | hp2 | a0001 | c0001 | t0032 | g0066 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02647 | hp1 | a0005 | c0003 | t0011 | g0018 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02647 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02698 | hp1 | a0002 | c0002 | t0009 | g0106 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02698 | hp2 | a0001 | c0001 | t0030 | g0004 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02717 | hp1 | a0001 | c0001 | t0026 | g0108 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02717 | hp2 | a0001 | c0001 | t0021 | g0116 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02723 | hp1 | a0002 | c0002 | t0003 | g0105 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02723 | hp2 | a0001 | c0001 | t0012 | g0019 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02735 | hp1 | a0001 | c0001 | t0002 | g0087 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02735 | hp2 | a0001 | c0001 | t0004 | g0005 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02738 | hp1 | a0001 | c0001 | t0004 | g0005 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02738 | hp2 | a0002 | c0002 | t0035 | g0095 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02809 | hp1 | a0001 | c0001 | t0001 | g0011 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02809 | hp2 | a0003 | c0005 | t0014 | g0010 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02818 | hp1 | a0001 | c0001 | t0012 | g0019 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02818 | hp2 | a0001 | c0001 | t0008 | g0007 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02886 | hp1 | a0002 | c0002 | t0003 | g0003 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02886 | hp2 | a0001 | c0001 | t0006 | g0047 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02896 | hp1 | a0001 | c0001 | t0047 | g0110 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02896 | hp2 | a0008 | c0010 | t0049 | g0120 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02922 | hp1 | a0001 | c0001 | t0001 | g0085 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02922 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02965 | hp1 | a0004 | c0004 | t0010 | g0020 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02965 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02970 | hp1 | a0004 | c0004 | t0010 | g0020 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02970 | hp2 | a0001 | c0001 | t0004 | g0005 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02976 | hp1 | a0002 | c0002 | t0003 | g0031 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02976 | hp2 | a0001 | c0001 | t0001 | g0073 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03017 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03017 | hp2 | a0001 | c0001 | t0002 | g0033 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03098 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03098 | hp2 | a0001 | c0001 | t0008 | g0060 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03130 | hp1 | a0001 | c0001 | t0001 | g0069 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03130 | hp2 | a0002 | c0002 | t0003 | g0031 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03139 | hp1 | a0001 | c0001 | t0004 | g0005 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03139 | hp2 | a0001 | c0001 | t0028 | g0005 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03195 | hp1 | a0001 | c0001 | t0020 | g0114 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03195 | hp2 | a0001 | c0001 | t0029 | g0045 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03209 | hp1 | a0001 | c0001 | t0024 | g0113 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03209 | hp2 | a0001 | c0001 | t0004 | g0005 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03225 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03225 | hp2 | a0001 | c0001 | t0008 | g0064 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03239 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03239 | hp2 | a0001 | c0001 | t0004 | g0005 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03453 | hp1 | a0005 | c0003 | t0011 | g0032 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03453 | hp2 | a0002 | c0002 | t0042 | g0017 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03486 | hp1 | a0005 | c0003 | t0011 | g0018 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03486 | hp2 | a0001 | c0001 | t0001 | g0016 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03491 | hp1 | a0001 | c0001 | t0002 | g0033 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03491 | hp2 | a0001 | c0001 | t0017 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03492 | hp1 | a0001 | c0001 | t0017 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03492 | hp2 | a0001 | c0001 | t0040 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03516 | hp1 | a0001 | c0007 | t0001 | g0089 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03516 | hp2 | a0001 | c0001 | t0001 | g0016 | AFR | ESN | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03540 | hp1 | a0001 | c0001 | t0019 | g0117 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03540 | hp2 | a0001 | c0001 | t0006 | g0002 | AFR | GWD | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03579 | hp1 | a0001 | c0001 | t0001 | g0011 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03579 | hp2 | a0004 | c0004 | t0010 | g0020 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03654 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03654 | hp2 | a0002 | c0002 | t0009 | g0003 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03669 | hp1 | a0001 | c0001 | t0033 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03669 | hp2 | a0001 | c0001 | t0006 | g0026 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03688 | hp1 | a0001 | c0001 | t0001 | g0015 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03688 | hp2 | a0002 | c0002 | t0003 | g0003 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03704 | hp1 | a0001 | c0001 | t0002 | g0001 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03704 | hp2 | a0002 | c0002 | t0003 | g0003 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03710 | hp1 | a0001 | c0001 | t0002 | g0002 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03710 | hp2 | a0001 | c0001 | t0002 | g0041 | SAS | PJL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03831 | hp1 | a0001 | c0001 | t0002 | g0048 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03831 | hp2 | a0001 | c0001 | t0002 | g0001 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03927 | hp1 | a0001 | c0001 | t0013 | g0092 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03927 | hp2 | a0001 | c0001 | t0001 | g0015 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03942 | hp1 | a0001 | c0008 | t0001 | g0070 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03942 | hp2 | a0001 | c0001 | t0002 | g0001 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04115 | hp1 | a0001 | c0001 | t0006 | g0026 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04115 | hp2 | a0001 | c0001 | t0001 | g0006 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04184 | hp1 | a0002 | c0002 | t0003 | g0003 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04184 | hp2 | a0001 | c0001 | t0001 | g0061 | SAS | BEB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04199 | hp1 | a0002 | c0002 | t0003 | g0003 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04199 | hp2 | a0001 | c0001 | t0002 | g0001 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04228 | hp1 | a0001 | c0001 | t0002 | g0054 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG04228 | hp2 | a0002 | c0002 | t0003 | g0090 | SAS | STU | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18522 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | YRI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18522 | hp2 | a0001 | c0001 | t0004 | g0005 | AFR | YRI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18612 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | CHB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18612 | hp2 | a0001 | c0001 | t0004 | g0005 | EAS | CHB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18747 | hp1 | a0001 | c0001 | t0001 | g0062 | EAS | CHB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18747 | hp2 | a0001 | c0001 | t0002 | g0079 | EAS | CHB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18906 | hp1 | a0003 | c0005 | t0014 | g0010 | AFR | YRI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18906 | hp2 | a0001 | c0001 | t0001 | g0013 | AFR | YRI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18943 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18943 | hp2 | a0002 | c0002 | t0003 | g0017 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18946 | hp1 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18946 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18947 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18947 | hp2 | a0001 | c0001 | t0005 | g0093 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18950 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18950 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18952 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18952 | hp2 | a0001 | c0001 | t0002 | g0024 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18954 | hp1 | a0001 | c0001 | t0006 | g0082 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18954 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18957 | hp1 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18957 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18961 | hp1 | a0001 | c0001 | t0001 | g0053 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18961 | hp2 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18963 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18963 | hp2 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18964 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18964 | hp2 | a0001 | c0001 | t0006 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18965 | hp1 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18965 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18967 | hp1 | a0001 | c0001 | t0007 | g0091 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18967 | hp2 | a0001 | c0001 | t0001 | g0112 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18971 | hp1 | a0001 | c0001 | t0034 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18971 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18972 | hp1 | a0001 | c0001 | t0002 | g0036 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18972 | hp2 | a0001 | c0001 | t0036 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18975 | hp1 | a0001 | c0001 | t0002 | g0086 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18975 | hp2 | a0002 | c0002 | t0003 | g0029 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18977 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18977 | hp2 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18978 | hp1 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18978 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18979 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18979 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18981 | hp1 | a0001 | c0001 | t0005 | g0027 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18981 | hp2 | a0001 | c0001 | t0002 | g0025 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18982 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18982 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18983 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18983 | hp2 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18984 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18984 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18986 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18986 | hp2 | a0001 | c0001 | t0001 | g0023 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18987 | hp1 | a0002 | c0002 | t0003 | g0102 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18987 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18993 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18993 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18995 | hp1 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18995 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18997 | hp1 | a0001 | c0001 | t0046 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18997 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18998 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18998 | hp2 | a0002 | c0002 | t0003 | g0100 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19000 | hp1 | a0001 | c0001 | t0005 | g0027 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19000 | hp2 | a0001 | c0001 | t0001 | g0049 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19001 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19001 | hp2 | a0002 | c0002 | t0003 | g0017 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19002 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19002 | hp2 | a0001 | c0001 | t0002 | g0057 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19005 | hp1 | a0001 | c0001 | t0001 | g0071 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19005 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19006 | hp1 | a0002 | c0002 | t0003 | g0101 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19006 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19007 | hp1 | a0001 | c0001 | t0006 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19007 | hp2 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19010 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19010 | hp2 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19011 | hp1 | a0001 | c0001 | t0006 | g0056 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19011 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19012 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19012 | hp2 | a0001 | c0001 | t0001 | g0006 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19030 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | LWK | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19030 | hp2 | a0001 | c0001 | t0001 | g0072 | AFR | LWK | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19043 | hp1 | a0001 | c0001 | t0039 | g0051 | AFR | LWK | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19043 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | LWK | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19055 | hp1 | a0001 | c0001 | t0007 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19055 | hp2 | a0001 | c0001 | t0006 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19056 | hp1 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19056 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19059 | hp1 | a0001 | c0001 | t0002 | g0077 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19059 | hp2 | a0001 | c0001 | t0025 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19060 | hp1 | a0002 | c0002 | t0003 | g0104 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19060 | hp2 | a0001 | c0001 | t0038 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19063 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19063 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19064 | hp1 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19064 | hp2 | a0002 | c0002 | t0003 | g0003 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19066 | hp1 | a0001 | c0001 | t0001 | g0006 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19066 | hp2 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19072 | hp1 | a0002 | c0002 | t0003 | g0030 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19072 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19076 | hp1 | a0001 | c0001 | t0007 | g0028 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19076 | hp2 | a0001 | c0001 | t0002 | g0001 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19084 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19084 | hp2 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19087 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19087 | hp2 | a0001 | c0001 | t0005 | g0094 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19091 | hp1 | a0001 | c0001 | t0006 | g0011 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19091 | hp2 | a0001 | c0001 | t0002 | g0037 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19240 | hp1 | a0001 | c0001 | t0008 | g0007 | AFR | YRI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA19240 | hp2 | a0003 | c0005 | t0023 | g0035 | AFR | YRI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20129 | hp1 | a0006 | c0006 | t0018 | g0001 | AFR | ASW | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20129 | hp2 | a0003 | c0005 | t0014 | g0010 | AFR | ASW | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20752 | hp1 | a0001 | c0001 | t0002 | g0001 | EUR | TSI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20752 | hp2 | a0002 | c0002 | t0009 | g0003 | EUR | TSI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20805 | hp1 | a0001 | c0001 | t0027 | g0005 | EUR | TSI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20805 | hp2 | a0001 | c0001 | t0037 | g0001 | EUR | TSI | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02109 | hp1 | a0002 | c0002 | t0003 | g0003 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02109 | hp2 | a0001 | c0001 | t0001 | g0065 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02559 | hp1 | a0001 | c0001 | t0045 | g0096 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG02559 | hp2 | a0001 | c0001 | t0048 | g0111 | AFR | ACB | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03471 | hp1 | a0005 | c0003 | t0011 | g0032 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG03471 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | MSL | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG06807 | hp1 | a0004 | c0004 | t0010 | g0119 | AFR | USA | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| HG06807 | hp2 | a0001 | c0001 | t0001 | g0050 | AFR | USA | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18955 | hp1 | a0001 | c0001 | t0002 | g0002 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA18955 | hp2 | a0001 | c0001 | t0005 | g0004 | EAS | JPT | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20300 | hp1 | a0001 | c0001 | t0001 | g0009 | AFR | USA | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA20300 | hp2 | a0001 | c0001 | t0001 | g0076 | AFR | USA | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA21309 | hp1 | a0001 | c0001 | t0001 | g0058 | AFR | LWK | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| NA21309 | hp2 | a0001 | c0001 | t0008 | g0081 | AFR | LWK | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| homoSapiens_chm13v2 | hp1 | a0002 | c0002 | t0009 | g0107 | REF | REF | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| homoSapiens_grch38 | hp1 | a0001 | c0001 | t0002 | g0001 | REF | REF | CFAP107_chr1_12741200_12768699 | CFAP107 | chr1 | 12741200 | 12768699 |
| chr:pos | ref | alt | # # of ahapid:amino-acid(protein) level |
ahapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:12746530
|
G | T | 1 | a0007 | 1 | HG01261.hp1 | stop_gained | HIGH | c.100G>T | p.Glu34* | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/4 | 331/3564 | 100/585 | 34/194 | chr1 | 12746530 | ||
| chr1:12755700
|
T | G | 1 | a0006 | 2 | HG00140.hp1 NA20129.hp1 |
missense_variant&splice_region_variant | MODERATE | c.112T>G | p.Phe38Val | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/4 | 343/3564 | 112/585 | 38/194 | chr1 | 12755700 | ||
| chr1:12755722
|
C | T | 1 | a0003 | 5 | HG00642.hp1 HG02809.hp2 NA18906.hp1 others(2): Show |
missense_variant | MODERATE | c.134C>T | p.Thr45Ile | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/4 | 365/3564 | 134/585 | 45/194 | chr1 | 12755722 | ||
| chr1:12759410
|
C | T | 1 | a0004 | 5 | HG02257.hp1 HG02965.hp1 HG02970.hp1 others(2): Show |
missense_variant | MODERATE | c.326C>T | p.Pro109Leu | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/4 | 557/3564 | 326/585 | 109/194 | chr1 | 12759410 | ||
| chr1:12760937
|
C | T | 1 | a0002 | 68 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(65): Show |
missense_variant | MODERATE | c.571C>T | p.Leu191Phe | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 802/3564 | 571/585 | 191/194 | chr1 | 12760937 | ||
| chr1:12760938
|
T | A | 1 | a0005 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
missense_variant | MODERATE | c.572T>A | p.Leu191His | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 803/3564 | 572/585 | 191/194 | chr1 | 12760938 |
| chr:pos | ref | alt | # # of chapid |
chapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:12755708
|
A | G | 1 | a0001c0008 | 1 | HG03942.hp1 | synonymous_variant | LOW | c.120A>G | p.Arg40Arg | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/4 | 351/3564 | 120/585 | 40/194 | chr1 | 12755708 | ||
| chr1:12759468
|
G | A | 1 | a0001c0007 | 1 | HG03516.hp1 | synonymous_variant | LOW | c.384G>A | p.Lys128Lys | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/4 | 615/3564 | 384/585 | 128/194 | chr1 | 12759468 |
| chr:pos | ref | alt | # # of thapid:transcript level |
thapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:12746225
|
G | A | 8 | a0001c0001t0012a0001c0001t0019a0001c0001t0020others(5): Show | 17 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(14): Show |
5_prime_UTR_variant | MODIFIER | c.-206G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/4 | 206 | chr1 | 12746225 | |||||
| chr1:12746230
|
A | C | 8 | a0001c0001t0012a0001c0001t0019a0001c0001t0020others(5): Show | 17 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(14): Show |
5_prime_UTR_variant | MODIFIER | c.-201A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/4 | 201 | chr1 | 12746230 | |||||
| chr1:12746248
|
C | A | 20 | a0001c0001t0004a0001c0001t0005a0001c0001t0012others(17): Show | 62 | HG00597.hp2 HG00642.hp1 HG01106.hp2 others(59): Show |
5_prime_UTR_variant | MODIFIER | c.-183C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/4 | 183 | chr1 | 12746248 | |||||
| chr1:12746293
|
T | A | 2 | a0001c0001t0024a0005c0003t0011 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
5_prime_UTR_variant | MODIFIER | c.-138T>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/4 | 138 | chr1 | 12746293 | |||||
| chr1:12746377
|
C | T | 3 | a0001c0001t0030a0001c0001t0047a0001c0001t0048 | 3 | HG02559.hp2 HG02698.hp2 HG02896.hp1 |
5_prime_UTR_premature_start_codon_gain_variant | LOW | c.-54C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/4 | chr1 | 12746377 | ||||||
| chr1:12760954
|
G | C | 1 | a0001c0001t0031 | 1 | HG00140.hp2 | 3_prime_UTR_variant | MODIFIER | c.*3G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 3 | chr1 | 12760954 | |||||
| chr1:12761142
|
A | G | 7 | a0001c0001t0004a0001c0001t0008a0001c0001t0016others(4): Show | 29 | HG01106.hp2 HG01167.hp2 HG01192.hp1 others(26): Show |
3_prime_UTR_variant | MODIFIER | c.*191A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 191 | chr1 | 12761142 | |||||
| chr1:12761294
|
C | T | 1 | a0001c0001t0026 | 1 | HG02717.hp1 | 3_prime_UTR_variant | MODIFIER | c.*343C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 343 | chr1 | 12761294 | |||||
| chr1:12761467
|
C | G | 1 | a0001c0001t0046 | 1 | NA18997.hp1 | 3_prime_UTR_variant | MODIFIER | c.*516C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 516 | chr1 | 12761467 | |||||
| chr1:12761487
|
A | G | 5 | a0001c0001t0005a0001c0001t0007a0001c0001t0013others(2): Show | 28 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(25): Show |
3_prime_UTR_variant | MODIFIER | c.*536A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 536 | chr1 | 12761487 | |||||
| chr1:12761519
|
G | A | 7 | a0001c0001t0004a0001c0001t0008a0001c0001t0016others(4): Show | 29 | HG01106.hp2 HG01167.hp2 HG01192.hp1 others(26): Show |
3_prime_UTR_variant | MODIFIER | c.*568G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 568 | chr1 | 12761519 | |||||
| chr1:12761543
|
G | GT | 19 | a0001c0001t0004a0001c0001t0005a0001c0001t0007others(16): Show | 71 | HG00280.hp1 HG00597.hp2 HG00642.hp1 others(68): Show |
3_prime_UTR_variant | MODIFIER | c.*603dupT | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 604 | INFO_REALIGN_3_PRIME | chr1 | 12761543 | ||||
| chr1:12761543
|
G | GTT | 4 | a0002c0002t0003a0002c0002t0009a0002c0002t0042others(1): Show | 67 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(64): Show |
3_prime_UTR_variant | MODIFIER | c.*602_*603dupTT | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 604 | INFO_REALIGN_3_PRIME | chr1 | 12761543 | ||||
| chr1:12761543
|
G | T | 4 | a0001c0001t0025a0001c0001t0029a0001c0001t0044others(1): Show | 4 | HG02451.hp1 HG02559.hp1 HG03195.hp2 others(1): Show |
3_prime_UTR_variant | MODIFIER | c.*592G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 592 | chr1 | 12761543 | |||||
| chr1:12761662
|
G | A | 1 | a0001c0001t0032 | 1 | HG02630.hp2 | 3_prime_UTR_variant | MODIFIER | c.*711G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 711 | chr1 | 12761662 | |||||
| chr1:12761683
|
C | A | 1 | a0005c0003t0011 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
3_prime_UTR_variant | MODIFIER | c.*732C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 732 | chr1 | 12761683 | |||||
| chr1:12761684
|
G | A | 4 | a0001c0001t0006a0001c0001t0017a0001c0001t0033others(1): Show | 17 | HG01192.hp2 HG02074.hp1 HG02155.hp1 others(14): Show |
3_prime_UTR_variant | MODIFIER | c.*733G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 733 | chr1 | 12761684 | |||||
| chr1:12761686
|
A | T | 1 | a0006c0006t0018 | 2 | HG00140.hp1 NA20129.hp1 |
3_prime_UTR_variant | MODIFIER | c.*735A>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 735 | chr1 | 12761686 | |||||
| chr1:12761693
|
C | T | 2 | a0003c0005t0022a0003c0005t0023 | 2 | HG00642.hp1 NA19240.hp2 |
3_prime_UTR_variant | MODIFIER | c.*742C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 742 | chr1 | 12761693 | |||||
| chr1:12761720
|
A | G | 2 | a0001c0001t0015a0001c0001t0041 | 4 | HG00280.hp1 HG01358.hp1 HG01515.hp1 others(1): Show |
3_prime_UTR_variant | MODIFIER | c.*769A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 769 | chr1 | 12761720 | |||||
| chr1:12761777
|
C | T | 1 | a0004c0004t0010 | 5 | HG02257.hp1 HG02965.hp1 HG02970.hp1 others(2): Show |
3_prime_UTR_variant | MODIFIER | c.*826C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 826 | chr1 | 12761777 | |||||
| chr1:12761787
|
G | A | 1 | a0001c0001t0017 | 2 | HG03491.hp2 HG03492.hp1 |
3_prime_UTR_variant | MODIFIER | c.*836G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 836 | chr1 | 12761787 | |||||
| chr1:12761816
|
G | A | 1 | a0001c0001t0034 | 1 | NA18971.hp1 | 3_prime_UTR_variant | MODIFIER | c.*865G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 865 | chr1 | 12761816 | |||||
| chr1:12761873
|
A | G | 1 | a0005c0003t0011 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
3_prime_UTR_variant | MODIFIER | c.*922A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 922 | chr1 | 12761873 | |||||
| chr1:12761875
|
C | A | 7 | a0001c0001t0019a0001c0001t0020a0001c0001t0021others(4): Show | 11 | HG00642.hp1 HG02257.hp1 HG02717.hp2 others(8): Show |
3_prime_UTR_variant | MODIFIER | c.*924C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 924 | chr1 | 12761875 | |||||
| chr1:12761938
|
C | T | 1 | a0001c0001t0040 | 1 | HG03492.hp2 | 3_prime_UTR_variant | MODIFIER | c.*987C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 987 | chr1 | 12761938 | |||||
| chr1:12761967
|
T | C | 1 | a0001c0001t0027 | 1 | NA20805.hp1 | 3_prime_UTR_variant | MODIFIER | c.*1016T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1016 | chr1 | 12761967 | |||||
| chr1:12762107
|
G | C | 1 | a0001c0001t0013 | 4 | HG01516.hp1 HG01517.hp1 HG02451.hp2 others(1): Show |
3_prime_UTR_variant | MODIFIER | c.*1156G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1156 | chr1 | 12762107 | |||||
| chr1:12762119
|
C | T | 1 | a0003c0005t0023 | 1 | NA19240.hp2 | 3_prime_UTR_variant | MODIFIER | c.*1168C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1168 | chr1 | 12762119 | |||||
| chr1:12762156
|
C | CCACCTAC others(206): Show |
6 | a0001c0001t0026a0001c0001t0044a0002c0002t0003others(3): Show | 69 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(66): Show |
3_prime_UTR_variant | MODIFIER | c.*1239_*1240insCTCC others(209): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1240 | INFO_REALIGN_3_PRIME | chr1 | 12762156 | ||||
| chr1:12762156
|
C | CCACCTAC others(418): Show |
2 | a0003c0005t0022a0003c0005t0023 | 2 | HG00642.hp1 NA19240.hp2 |
3_prime_UTR_variant | MODIFIER | c.*1239_*1240insCTCC others(421): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1240 | INFO_REALIGN_3_PRIME | chr1 | 12762156 | ||||
| chr1:12762156
|
C | CCACCTAC others(206): Show |
1 | a0001c0001t0036 | 1 | NA18972.hp2 | 3_prime_UTR_variant | MODIFIER | c.*1346_*1558dupAATG others(209): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1559 | INFO_REALIGN_3_PRIME | chr1 | 12762156 | ||||
| chr1:12762191
|
T | C | 23 | a0001c0001t0004a0001c0001t0005a0001c0001t0007others(20): Show | 79 | HG00544.hp1 HG00597.hp2 HG01106.hp2 others(76): Show |
3_prime_UTR_variant | MODIFIER | c.*1240T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1240 | chr1 | 12762191 | |||||
| chr1:12762226
|
C | T | 7 | a0001c0001t0044a0002c0002t0003a0002c0002t0009others(4): Show | 70 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(67): Show |
3_prime_UTR_variant | MODIFIER | c.*1275C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1275 | chr1 | 12762226 | |||||
| chr1:12762227
|
G | A | 21 | a0001c0001t0004a0001c0001t0005a0001c0001t0007others(18): Show | 73 | HG00544.hp1 HG00597.hp2 HG01106.hp2 others(70): Show |
3_prime_UTR_variant | MODIFIER | c.*1276G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1276 | chr1 | 12762227 | |||||
| chr1:12762280
|
T | C | 1 | a0002c0002t0009 | 6 | HG01070.hp2 HG01255.hp2 HG02698.hp1 others(3): Show |
3_prime_UTR_variant | MODIFIER | c.*1329T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1329 | chr1 | 12762280 | |||||
| chr1:12762350
|
CA | C | 17 | a0001c0001t0004a0001c0001t0008a0001c0001t0016others(14): Show | 106 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(103): Show |
3_prime_UTR_variant | MODIFIER | c.*1400delA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1400 | chr1 | 12762350 | |||||
| chr1:12762351
|
A | AGGGGGTG others(417): Show |
1 | a0002c0002t0043 | 1 | HG00544.hp1 | 3_prime_UTR_variant | MODIFIER | c.*1487_*1488insTGCT others(420): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1488 | INFO_REALIGN_3_PRIME | chr1 | 12762351 | ||||
| chr1:12762393
|
C | A | 1 | a0001c0001t0028 | 1 | HG03139.hp2 | 3_prime_UTR_variant | MODIFIER | c.*1442C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1442 | chr1 | 12762393 | |||||
| chr1:12762439
|
C | T | 20 | a0001c0001t0004a0001c0001t0008a0001c0001t0016others(17): Show | 113 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(110): Show |
3_prime_UTR_variant | MODIFIER | c.*1488C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1488 | chr1 | 12762439 | |||||
| chr1:12762518
|
G | A | 1 | a0001c0001t0033 | 1 | HG03669.hp1 | 3_prime_UTR_variant | MODIFIER | c.*1567G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1567 | chr1 | 12762518 | |||||
| chr1:12762567
|
G | A | 1 | a0001c0001t0028 | 1 | HG03139.hp2 | 3_prime_UTR_variant | MODIFIER | c.*1616G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1616 | chr1 | 12762567 | |||||
| chr1:12762835
|
A | G | 1 | a0001c0001t0039 | 1 | NA19043.hp1 | 3_prime_UTR_variant | MODIFIER | c.*1884A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1884 | chr1 | 12762835 | |||||
| chr1:12762889
|
T | TA | 46 | a0001c0001t0001a0001c0001t0004a0001c0001t0005others(43): Show | 263 | HG00140.hp1 HG00140.hp2 HG00280.hp1 others(260): Show |
3_prime_UTR_variant | MODIFIER | c.*1945dupA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 1946 | INFO_REALIGN_3_PRIME | chr1 | 12762889 | ||||
| chr1:12762953
|
T | C | 1 | a0001c0001t0037 | 1 | NA20805.hp2 | 3_prime_UTR_variant | MODIFIER | c.*2002T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2002 | chr1 | 12762953 | |||||
| chr1:12763016
|
G | A | 2 | a0001c0001t0047a0001c0001t0048 | 2 | HG02559.hp2 HG02896.hp1 |
3_prime_UTR_variant | MODIFIER | c.*2065G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2065 | chr1 | 12763016 | |||||
| chr1:12763073
|
C | T | 2 | a0001c0001t0016a0001c0001t0028 | 3 | HG01106.hp2 HG01192.hp1 HG03139.hp2 |
3_prime_UTR_variant | MODIFIER | c.*2122C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2122 | chr1 | 12763073 | |||||
| chr1:12763166
|
A | G | 2 | a0001c0001t0005a0001c0001t0025 | 14 | HG00597.hp2 HG02071.hp1 NA18946.hp1 others(11): Show |
3_prime_UTR_variant | MODIFIER | c.*2215A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2215 | chr1 | 12763166 | |||||
| chr1:12763249
|
C | T | 9 | a0001c0001t0005a0001c0001t0007a0001c0001t0013others(6): Show | 36 | HG00597.hp2 HG00642.hp1 HG01516.hp1 others(33): Show |
3_prime_UTR_variant | MODIFIER | c.*2298C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2298 | chr1 | 12763249 | |||||
| chr1:12763473
|
G | A | 1 | a0001c0001t0045 | 1 | HG02559.hp1 | 3_prime_UTR_variant | MODIFIER | c.*2522G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2522 | chr1 | 12763473 | |||||
| chr1:12763475
|
G | A | 1 | a0001c0001t0047 | 1 | HG02896.hp1 | 3_prime_UTR_variant | MODIFIER | c.*2524G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2524 | chr1 | 12763475 | |||||
| chr1:12763488
|
G | A | 7 | a0001c0001t0005a0001c0001t0007a0001c0001t0013others(4): Show | 30 | HG00597.hp2 HG00642.hp1 HG01516.hp1 others(27): Show |
3_prime_UTR_variant | MODIFIER | c.*2537G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2537 | chr1 | 12763488 | |||||
| chr1:12763519
|
A | G | 28 | a0001c0001t0004a0001c0001t0005a0001c0001t0007others(25): Show | 144 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(141): Show |
3_prime_UTR_variant | MODIFIER | c.*2568A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2568 | chr1 | 12763519 | |||||
| chr1:12763523
|
G | A | 1 | a0002c0002t0042 | 1 | HG03453.hp2 | 3_prime_UTR_variant | MODIFIER | c.*2572G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2572 | chr1 | 12763523 | |||||
| chr1:12763608
|
A | G | 1 | a0001c0001t0038 | 1 | NA19060.hp2 | 3_prime_UTR_variant | MODIFIER | c.*2657A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 4/4 | 2657 | chr1 | 12763608 |
| chr:pos | ref | alt | # # of ghapid:genebody level |
ghapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | genebody_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr1:12746586
|
T | C | 3 | a0003c0005t0014g0010a0003c0005t0022g0034a0003c0005t0023g0035 | 5 | HG00642.hp1 HG02809.hp2 NA18906.hp1 others(2): Show |
intron_variant | MODIFIER | c.111+45T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12746586 | ||||||
| chr1:12746764
|
C | T | 1 | a0001c0001t0002g0033 | 2 | HG03017.hp2 HG03491.hp1 |
intron_variant | MODIFIER | c.111+223C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12746764 | ||||||
| chr1:12746795
|
T | C | 1 | a0001c0001t0002g0036 | 1 | NA18972.hp1 | intron_variant | MODIFIER | c.111+254T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12746795 | ||||||
| chr1:12746879
|
T | C | 1 | a0001c0001t0002g0037 | 1 | NA19091.hp2 | intron_variant | MODIFIER | c.111+338T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12746879 | ||||||
| chr1:12747071
|
C | CT | 4 | a0001c0001t0001g0112a0001c0001t0024g0113a0005c0003t0011g0018others(1): Show | 7 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(4): Show |
intron_variant | MODIFIER | c.111+544dupT | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12747071 | |||||
| chr1:12747071
|
C | CTT | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+543_111+544dup others(2): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12747071 | |||||
| chr1:12747090
|
C | A | 1 | a0001c0001t0020g0114 | 1 | HG03195.hp1 | intron_variant | MODIFIER | c.111+549C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12747090 | ||||||
| chr1:12747272
|
T | C | 1 | a0001c0001t0001g0038 | 1 | HG02165.hp2 | intron_variant | MODIFIER | c.111+731T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12747272 | ||||||
| chr1:12747329
|
C | A | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+788C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12747329 | ||||||
| chr1:12747353
|
G | A | 6 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0016g0005others(3): Show | 17 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(14): Show |
intron_variant | MODIFIER | c.111+812G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12747353 | ||||||
| chr1:12747467
|
G | T | 2 | a0001c0001t0047g0110a0001c0001t0048g0111 | 2 | HG02559.hp2 HG02896.hp1 |
intron_variant | MODIFIER | c.111+926G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12747467 | ||||||
| chr1:12747927
|
G | A | 3 | a0001c0001t0002g0021a0001c0001t0002g0041a0001c0001t0002g0042 | 4 | HG01361.hp1 HG01978.hp1 HG02300.hp1 others(1): Show |
intron_variant | MODIFIER | c.111+1386G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12747927 | ||||||
| chr1:12748244
|
G | T | 1 | a0001c0001t0044g0109 | 1 | HG02451.hp1 | intron_variant | MODIFIER | c.111+1703G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748244 | ||||||
| chr1:12748253
|
G | C | 3 | a0002c0002t0003g0008a0002c0002t0003g0043a0002c0002t0003g0044 | 6 | HG00323.hp2 HG00639.hp1 HG01069.hp1 others(3): Show |
intron_variant | MODIFIER | c.111+1712G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748253 | ||||||
| chr1:12748389
|
G | A | 8 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(5): Show | 19 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(16): Show |
intron_variant | MODIFIER | c.111+1848G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748389 | ||||||
| chr1:12748461
|
T | TA | 11 | a0001c0001t0001g0011a0001c0001t0001g0049a0001c0001t0001g0050others(8): Show | 12 | HG00741.hp2 HG02451.hp1 HG02809.hp1 others(9): Show |
intron_variant | MODIFIER | c.111+1939dupA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12748461 | |||||
| chr1:12748474
|
A | AC | 13 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(10): Show | 29 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(26): Show |
intron_variant | MODIFIER | c.111+1933_111+1934i others(3): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748474 | ||||||
| chr1:12748474
|
A | C | 48 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(45): Show | 111 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(108): Show |
intron_variant | MODIFIER | c.111+1933A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748474 | ||||||
| chr1:12748489
|
A | T | 1 | a0001c0007t0001g0089 | 1 | HG03516.hp1 | intron_variant | MODIFIER | c.111+1948A>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748489 | ||||||
| chr1:12748519
|
G | T | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.111+1978G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748519 | ||||||
| chr1:12748543
|
G | A | 1 | a0001c0001t0016g0039 | 1 | HG01106.hp2 | intron_variant | MODIFIER | c.111+2002G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748543 | ||||||
| chr1:12748556
|
C | T | 1 | a0001c0001t0001g0088 | 1 | HG02258.hp2 | intron_variant | MODIFIER | c.111+2015C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748556 | ||||||
| chr1:12748580
|
G | T | 3 | a0003c0005t0014g0010a0003c0005t0022g0034a0003c0005t0023g0035 | 5 | HG00642.hp1 HG02809.hp2 NA18906.hp1 others(2): Show |
intron_variant | MODIFIER | c.111+2039G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748580 | ||||||
| chr1:12748643
|
A | G | 24 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(21): Show | 46 | HG00597.hp2 HG00642.hp1 HG01109.hp1 others(43): Show |
intron_variant | MODIFIER | c.111+2102A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748643 | ||||||
| chr1:12748660
|
C | T | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+2119C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748660 | ||||||
| chr1:12748689
|
GA | G | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+2157delA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12748689 | |||||
| chr1:12748725
|
AGT | A | 3 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0021g0116 | 5 | HG01109.hp1 HG02280.hp2 HG02717.hp2 others(2): Show |
intron_variant | MODIFIER | c.111+2186_111+2187d others(4): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12748725 | |||||
| chr1:12748864
|
A | G | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.111+2323A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748864 | ||||||
| chr1:12748877
|
A | G | 2 | a0001c0001t0047g0110a0001c0001t0048g0111 | 2 | HG02559.hp2 HG02896.hp1 |
intron_variant | MODIFIER | c.111+2336A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748877 | ||||||
| chr1:12748996
|
A | G | 3 | a0003c0005t0014g0010a0003c0005t0022g0034a0003c0005t0023g0035 | 5 | HG00642.hp1 HG02809.hp2 NA18906.hp1 others(2): Show |
intron_variant | MODIFIER | c.111+2455A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12748996 | ||||||
| chr1:12749032
|
C | A | 1 | a0002c0002t0035g0095 | 1 | HG02738.hp2 | intron_variant | MODIFIER | c.111+2491C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749032 | ||||||
| chr1:12749096
|
T | C | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+2555T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749096 | ||||||
| chr1:12749235
|
A | C | 1 | a0001c0001t0002g0054 | 1 | HG04228.hp1 | intron_variant | MODIFIER | c.111+2694A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749235 | ||||||
| chr1:12749245
|
C | T | 1 | a0003c0005t0023g0035 | 1 | NA19240.hp2 | intron_variant | MODIFIER | c.111+2704C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749245 | ||||||
| chr1:12749325
|
T | C | 1 | a0001c0001t0045g0096 | 1 | HG02559.hp1 | intron_variant | MODIFIER | c.111+2784T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749325 | ||||||
| chr1:12749334
|
C | G | 1 | a0002c0002t0009g0107 | 1 | homoSapiens_chm13v2.hp1 | intron_variant | MODIFIER | c.111+2793C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749334 | ||||||
| chr1:12749390
|
T | C | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.111+2849T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749390 | ||||||
| chr1:12749527
|
C | T | 3 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0021g0116 | 5 | HG01109.hp1 HG02280.hp2 HG02717.hp2 others(2): Show |
intron_variant | MODIFIER | c.111+2986C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749527 | ||||||
| chr1:12749592
|
C | G | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+3051C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749592 | ||||||
| chr1:12749652
|
G | GA | 36 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(33): Show | 72 | HG00597.hp2 HG00642.hp1 HG01106.hp2 others(69): Show |
intron_variant | MODIFIER | c.111+3112dupA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12749652 | |||||
| chr1:12749700
|
A | G | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.111+3159A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749700 | ||||||
| chr1:12749753
|
A | C | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.111+3212A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749753 | ||||||
| chr1:12749926
|
A | T | 12 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(9): Show | 28 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(25): Show |
intron_variant | MODIFIER | c.111+3385A>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749926 | ||||||
| chr1:12749951
|
T | G | 1 | a0001c0001t0001g0055 | 1 | HG01256.hp2 | intron_variant | MODIFIER | c.111+3410T>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749951 | ||||||
| chr1:12749979
|
G | T | 1 | a0001c0001t0024g0113 | 1 | HG03209.hp1 | intron_variant | MODIFIER | c.111+3438G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12749979 | ||||||
| chr1:12750044
|
G | C | 1 | a0001c0001t0006g0056 | 1 | NA19011.hp1 | intron_variant | MODIFIER | c.111+3503G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750044 | ||||||
| chr1:12750148
|
T | C | 1 | a0001c0001t0006g0047 | 1 | HG02886.hp2 | intron_variant | MODIFIER | c.111+3607T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750148 | ||||||
| chr1:12750188
|
T | C | 1 | a0001c0001t0002g0057 | 1 | NA19002.hp2 | intron_variant | MODIFIER | c.111+3647T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750188 | ||||||
| chr1:12750230
|
A | G | 24 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(21): Show | 46 | HG00597.hp2 HG00642.hp1 HG01109.hp1 others(43): Show |
intron_variant | MODIFIER | c.111+3689A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750230 | ||||||
| chr1:12750241
|
A | G | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.111+3700A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750241 | ||||||
| chr1:12750272
|
C | A | 3 | a0002c0002t0003g0097a0002c0002t0003g0098a0002c0002t0003g0099 | 3 | HG00323.hp1 HG01175.hp2 HG01358.hp2 |
intron_variant | MODIFIER | c.111+3731C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750272 | ||||||
| chr1:12750380
|
A | G | 12 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(9): Show | 28 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(25): Show |
intron_variant | MODIFIER | c.111+3839A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750380 | ||||||
| chr1:12750620
|
G | A | 9 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(6): Show | 13 | HG01109.hp1 HG02257.hp1 HG02280.hp2 others(10): Show |
intron_variant | MODIFIER | c.111+4079G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750620 | ||||||
| chr1:12750639
|
T | C | 1 | a0001c0001t0029g0045 | 1 | HG03195.hp2 | intron_variant | MODIFIER | c.111+4098T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750639 | ||||||
| chr1:12750930
|
A | G | 1 | a0001c0001t0001g0009 | 4 | HG00639.hp2 HG00738.hp1 HG00741.hp1 others(1): Show |
intron_variant | MODIFIER | c.111+4389A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750930 | ||||||
| chr1:12750980
|
C | T | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.111+4439C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12750980 | ||||||
| chr1:12751002
|
T | C | 1 | a0001c0001t0002g0012 | 3 | HG00735.hp1 HG01256.hp1 HG01258.hp1 |
intron_variant | MODIFIER | c.111+4461T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751002 | ||||||
| chr1:12751087
|
T | C | 1 | a0002c0002t0003g0043 | 1 | HG00639.hp1 | intron_variant | MODIFIER | c.111+4546T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751087 | ||||||
| chr1:12751140
|
T | G | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.112-4560T>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751140 | ||||||
| chr1:12751144
|
A | G | 1 | a0002c0002t0009g0106 | 1 | HG02698.hp1 | intron_variant | MODIFIER | c.112-4556A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751144 | ||||||
| chr1:12751196
|
G | A | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-4504G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751196 | ||||||
| chr1:12751326
|
A | G | 14 | a0001c0001t0002g0033a0001c0001t0002g0087a0001c0001t0005g0004others(11): Show | 31 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(28): Show |
intron_variant | MODIFIER | c.112-4374A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751326 | ||||||
| chr1:12751353
|
G | A | 8 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(5): Show | 19 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(16): Show |
intron_variant | MODIFIER | c.112-4347G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751353 | ||||||
| chr1:12751419
|
G | A | 1 | a0001c0001t0002g0048 | 1 | HG03831.hp1 | intron_variant | MODIFIER | c.112-4281G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751419 | ||||||
| chr1:12751456
|
G | A | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-4244G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751456 | ||||||
| chr1:12751490
|
C | T | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-4210C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751490 | ||||||
| chr1:12751613
|
C | T | 1 | a0001c0001t0045g0096 | 1 | HG02559.hp1 | intron_variant | MODIFIER | c.112-4087C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751613 | ||||||
| chr1:12751638
|
G | A | 1 | a0001c0001t0001g0058 | 1 | NA21309.hp1 | intron_variant | MODIFIER | c.112-4062G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751638 | ||||||
| chr1:12751701
|
A | G | 12 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(9): Show | 28 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(25): Show |
intron_variant | MODIFIER | c.112-3999A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751701 | ||||||
| chr1:12751723
|
A | C | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.112-3977A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751723 | ||||||
| chr1:12751859
|
C | T | 3 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0021g0116 | 5 | HG01109.hp1 HG02280.hp2 HG02717.hp2 others(2): Show |
intron_variant | MODIFIER | c.112-3841C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751859 | ||||||
| chr1:12751861
|
T | C | 1 | a0001c0001t0001g0059 | 1 | HG02055.hp1 | intron_variant | MODIFIER | c.112-3839T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751861 | ||||||
| chr1:12751944
|
G | C | 1 | a0002c0002t0003g0100 | 1 | NA18998.hp2 | intron_variant | MODIFIER | c.112-3756G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751944 | ||||||
| chr1:12751961
|
G | A | 1 | a0005c0003t0011g0018 | 3 | HG02055.hp2 HG02647.hp1 HG03486.hp1 |
intron_variant | MODIFIER | c.112-3739G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751961 | ||||||
| chr1:12751961
|
G | C | 1 | a0001c0001t0001g0013 | 3 | HG02615.hp2 HG02622.hp1 NA18906.hp2 |
intron_variant | MODIFIER | c.112-3739G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12751961 | ||||||
| chr1:12752140
|
G | A | 13 | a0001c0001t0001g0014a0001c0001t0012g0019a0001c0001t0012g0115others(10): Show | 21 | HG00642.hp1 HG01109.hp1 HG01928.hp1 others(18): Show |
intron_variant | MODIFIER | c.112-3560G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752140 | ||||||
| chr1:12752254
|
G | T | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-3446G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752254 | ||||||
| chr1:12752313
|
A | G | 24 | a0002c0002t0003g0003a0002c0002t0003g0008a0002c0002t0003g0017others(21): Show | 67 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(64): Show |
intron_variant | MODIFIER | c.112-3387A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752313 | ||||||
| chr1:12752325
|
C | G | 1 | a0001c0001t0002g0042 | 1 | HG01361.hp1 | intron_variant | MODIFIER | c.112-3375C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752325 | ||||||
| chr1:12752363
|
TC | T | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-3336delC | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752363 | ||||||
| chr1:12752413
|
T | C | 11 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(8): Show | 17 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(14): Show |
intron_variant | MODIFIER | c.112-3287T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752413 | ||||||
| chr1:12752435
|
A | C | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.112-3265A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752435 | ||||||
| chr1:12752445
|
C | T | 1 | a0001c0001t0006g0026 | 2 | HG03669.hp2 HG04115.hp1 |
intron_variant | MODIFIER | c.112-3255C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752445 | ||||||
| chr1:12752466
|
T | C | 3 | a0003c0005t0014g0010a0003c0005t0022g0034a0003c0005t0023g0035 | 5 | HG00642.hp1 HG02809.hp2 NA18906.hp1 others(2): Show |
intron_variant | MODIFIER | c.112-3234T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752466 | ||||||
| chr1:12752515
|
C | G | 1 | a0001c0001t0045g0096 | 1 | HG02559.hp1 | intron_variant | MODIFIER | c.112-3185C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752515 | ||||||
| chr1:12752545
|
C | T | 1 | a0003c0005t0022g0034 | 1 | HG00642.hp1 | intron_variant | MODIFIER | c.112-3155C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752545 | ||||||
| chr1:12752550
|
CTG | C | 2 | a0001c0001t0005g0027a0001c0001t0007g0091 | 3 | NA18967.hp1 NA18981.hp1 NA19000.hp1 |
intron_variant | MODIFIER | c.112-3148_112-3147d others(4): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12752550 | |||||
| chr1:12752552
|
G | C | 10 | a0001c0001t0005g0004a0001c0001t0005g0093a0001c0001t0005g0094others(7): Show | 25 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(22): Show |
intron_variant | MODIFIER | c.112-3148G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752552 | ||||||
| chr1:12752553
|
TCTA | T | 8 | a0001c0001t0005g0004a0001c0001t0005g0093a0001c0001t0005g0094others(5): Show | 23 | HG00597.hp2 HG01517.hp1 HG02071.hp1 others(20): Show |
intron_variant | MODIFIER | c.112-3146_112-3144d others(5): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752553 | ||||||
| chr1:12752555
|
T | TA | 54 | a0001c0001t0001g0002a0001c0001t0001g0007a0001c0001t0001g0009others(51): Show | 109 | HG00140.hp2 HG00323.hp1 HG00639.hp1 others(106): Show |
intron_variant | MODIFIER | c.112-3118dupA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12752555 | |||||
| chr1:12752555
|
T | TAA | 13 | a0001c0001t0001g0006a0001c0001t0001g0049a0001c0001t0001g0059others(10): Show | 21 | HG01109.hp1 HG01358.hp1 HG02055.hp1 others(18): Show |
intron_variant | MODIFIER | c.112-3119_112-3118d others(4): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12752555 | |||||
| chr1:12752555
|
TA | T | 11 | a0001c0001t0002g0025a0001c0001t0002g0087a0001c0001t0004g0005others(8): Show | 26 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(23): Show |
intron_variant | MODIFIER | c.112-3118delA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12752555 | |||||
| chr1:12752557
|
A | T | 3 | a0001c0001t0024g0113a0005c0003t0011g0018a0005c0003t0011g0032 | 6 | HG02055.hp2 HG02647.hp1 HG03209.hp1 others(3): Show |
intron_variant | MODIFIER | c.112-3143A>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752557 | ||||||
| chr1:12752850
|
G | A | 8 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(5): Show | 19 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(16): Show |
intron_variant | MODIFIER | c.112-2850G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752850 | ||||||
| chr1:12752956
|
G | A | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.112-2744G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12752956 | ||||||
| chr1:12753019
|
G | A | 1 | a0001c0001t0002g0075 | 1 | HG01981.hp1 | intron_variant | MODIFIER | c.112-2681G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753019 | ||||||
| chr1:12753083
|
A | G | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.112-2617A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753083 | ||||||
| chr1:12753115
|
A | G | 1 | a0001c0001t0006g0047 | 1 | HG02886.hp2 | intron_variant | MODIFIER | c.112-2585A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753115 | ||||||
| chr1:12753160
|
GGAGAT | G | 2 | a0001c0001t0001g0023a0001c0001t0001g0112 | 3 | HG02523.hp1 NA18967.hp2 NA18986.hp2 |
intron_variant | MODIFIER | c.112-2537_112-2533d others(7): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12753160 | |||||
| chr1:12753192
|
C | T | 1 | a0001c0001t0001g0058 | 1 | NA21309.hp1 | intron_variant | MODIFIER | c.112-2508C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753192 | ||||||
| chr1:12753312
|
A | G | 1 | a0001c0001t0002g0036 | 1 | NA18972.hp1 | intron_variant | MODIFIER | c.112-2388A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753312 | ||||||
| chr1:12753325
|
C | G | 8 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(5): Show | 19 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(16): Show |
intron_variant | MODIFIER | c.112-2375C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753325 | ||||||
| chr1:12753341
|
G | GT | 17 | a0001c0001t0001g0016a0001c0001t0001g0049a0001c0001t0001g0053others(14): Show | 24 | HG01109.hp1 HG01928.hp2 HG02257.hp1 others(21): Show |
intron_variant | MODIFIER | c.112-2347dupT | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12753341 | |||||
| chr1:12753342
|
T | G | 1 | a0001c0001t0045g0096 | 1 | HG02559.hp1 | intron_variant | MODIFIER | c.112-2358T>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753342 | ||||||
| chr1:12753348
|
T | C | 1 | a0002c0002t0003g0030 | 2 | HG00438.hp1 NA19072.hp1 |
intron_variant | MODIFIER | c.112-2352T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753348 | ||||||
| chr1:12753407
|
G | A | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-2293G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753407 | ||||||
| chr1:12753532
|
T | C | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-2168T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753532 | ||||||
| chr1:12753573
|
C | T | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-2127C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753573 | ||||||
| chr1:12753643
|
G | A | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.112-2057G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753643 | ||||||
| chr1:12753736
|
C | A | 1 | a0001c0001t0001g0062 | 1 | NA18747.hp1 | intron_variant | MODIFIER | c.112-1964C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753736 | ||||||
| chr1:12753770
|
A | G | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-1930A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753770 | ||||||
| chr1:12753902
|
C | T | 12 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(9): Show | 18 | HG00642.hp1 HG01109.hp1 HG02257.hp1 others(15): Show |
intron_variant | MODIFIER | c.112-1798C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753902 | ||||||
| chr1:12753952
|
A | T | 1 | a0001c0001t0001g0073 | 1 | HG02976.hp2 | intron_variant | MODIFIER | c.112-1748A>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12753952 | ||||||
| chr1:12754063
|
G | A | 1 | a0001c0001t0001g0076 | 1 | NA20300.hp2 | intron_variant | MODIFIER | c.112-1637G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754063 | ||||||
| chr1:12754099
|
A | G | 33 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(30): Show | 66 | HG00597.hp2 HG00642.hp1 HG01106.hp2 others(63): Show |
intron_variant | MODIFIER | c.112-1601A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754099 | ||||||
| chr1:12754285
|
A | C | 3 | a0003c0005t0014g0010a0003c0005t0022g0034a0003c0005t0023g0035 | 5 | HG00642.hp1 HG02809.hp2 NA18906.hp1 others(2): Show |
intron_variant | MODIFIER | c.112-1415A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754285 | ||||||
| chr1:12754307
|
A | G | 24 | a0002c0002t0003g0003a0002c0002t0003g0008a0002c0002t0003g0017others(21): Show | 67 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(64): Show |
intron_variant | MODIFIER | c.112-1393A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754307 | ||||||
| chr1:12754319
|
A | G | 8 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(5): Show | 12 | HG01109.hp1 HG02257.hp1 HG02280.hp2 others(9): Show |
intron_variant | MODIFIER | c.112-1381A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754319 | ||||||
| chr1:12754348
|
CTAGTATC others(38): Show |
C | 1 | a0001c0001t0002g0077 | 1 | NA19059.hp1 | intron_variant | MODIFIER | c.112-1351_112-1307d others(47): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754348 | ||||||
| chr1:12754456
|
A | C | 1 | a0002c0002t0003g0031 | 2 | HG02976.hp1 HG03130.hp2 |
intron_variant | MODIFIER | c.112-1244A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754456 | ||||||
| chr1:12754467
|
G | A | 2 | a0001c0001t0001g0063a0001c0001t0001g0074 | 2 | HG00735.hp2 HG01928.hp2 |
intron_variant | MODIFIER | c.112-1233G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754467 | ||||||
| chr1:12754512
|
T | C | 1 | a0001c0001t0001g0050 | 1 | HG06807.hp2 | intron_variant | MODIFIER | c.112-1188T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754512 | ||||||
| chr1:12754635
|
C | T | 1 | a0001c0001t0019g0117 | 1 | HG03540.hp1 | intron_variant | MODIFIER | c.112-1065C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754635 | ||||||
| chr1:12754710
|
A | G | 1 | a0001c0001t0001g0072 | 1 | NA19030.hp2 | intron_variant | MODIFIER | c.112-990A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754710 | ||||||
| chr1:12754954
|
C | T | 1 | a0001c0001t0001g0071 | 1 | NA19005.hp1 | intron_variant | MODIFIER | c.112-746C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754954 | ||||||
| chr1:12754970
|
C | A | 1 | a0001c0001t0008g0064 | 1 | HG03225.hp2 | intron_variant | MODIFIER | c.112-730C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12754970 | ||||||
| chr1:12755063
|
A | C | 1 | a0001c0001t0002g0054 | 1 | HG04228.hp1 | intron_variant | MODIFIER | c.112-637A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12755063 | ||||||
| chr1:12755175
|
G | C | 1 | a0001c0001t0001g0014 | 3 | HG01928.hp1 HG01934.hp1 HG02148.hp2 |
intron_variant | MODIFIER | c.112-525G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12755175 | ||||||
| chr1:12755340
|
G | C | 1 | a0001c0001t0001g0065 | 1 | HG02109.hp2 | intron_variant | MODIFIER | c.112-360G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12755340 | ||||||
| chr1:12755404
|
C | CA | 50 | a0001c0001t0002g0024a0001c0001t0002g0077a0001c0001t0004g0005others(47): Show | 114 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(111): Show |
intron_variant | MODIFIER | c.112-285dupA | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12755404 | |||||
| chr1:12755404
|
C | CAA | 15 | a0001c0001t0004g0040a0001c0001t0005g0004a0001c0001t0005g0027others(12): Show | 31 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(28): Show |
intron_variant | MODIFIER | c.112-286_112-285dup others(2): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | INFO_REALIGN_3_PRIME | chr1 | 12755404 | |||||
| chr1:12755570
|
G | A | 1 | a0003c0005t0023g0035 | 1 | NA19240.hp2 | intron_variant | MODIFIER | c.112-130G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12755570 | ||||||
| chr1:12755696
|
C | G | 2 | a0001c0001t0047g0110a0001c0001t0048g0111 | 2 | HG02559.hp2 HG02896.hp1 |
splice_region_variant&intron_variant | LOW | c.112-4C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 1/3 | chr1 | 12755696 | ||||||
| chr1:12755877
|
G | A | 12 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(9): Show | 28 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(25): Show |
intron_variant | MODIFIER | c.225+64G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12755877 | ||||||
| chr1:12755988
|
G | A | 1 | a0002c0002t0003g0098 | 1 | HG01175.hp2 | intron_variant | MODIFIER | c.225+175G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12755988 | ||||||
| chr1:12756005
|
T | C | 1 | a0001c0001t0002g0067 | 1 | HG02074.hp2 | intron_variant | MODIFIER | c.225+192T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756005 | ||||||
| chr1:12756270
|
G | A | 1 | a0001c0001t0002g0012 | 3 | HG00735.hp1 HG01256.hp1 HG01258.hp1 |
intron_variant | MODIFIER | c.225+457G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756270 | ||||||
| chr1:12756348
|
G | A | 24 | a0002c0002t0003g0003a0002c0002t0003g0008a0002c0002t0003g0017others(21): Show | 67 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(64): Show |
intron_variant | MODIFIER | c.225+535G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756348 | ||||||
| chr1:12756352
|
T | C | 1 | a0002c0002t0003g0098 | 1 | HG01175.hp2 | intron_variant | MODIFIER | c.225+539T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756352 | ||||||
| chr1:12756366
|
T | C | 53 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(50): Show | 121 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(118): Show |
intron_variant | MODIFIER | c.225+553T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756366 | ||||||
| chr1:12756416
|
G | T | 24 | a0002c0002t0003g0003a0002c0002t0003g0008a0002c0002t0003g0017others(21): Show | 67 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(64): Show |
intron_variant | MODIFIER | c.225+603G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756416 | ||||||
| chr1:12756421
|
C | T | 24 | a0002c0002t0003g0003a0002c0002t0003g0008a0002c0002t0003g0017others(21): Show | 67 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(64): Show |
intron_variant | MODIFIER | c.225+608C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756421 | ||||||
| chr1:12756465
|
C | T | 2 | a0005c0003t0011g0018a0005c0003t0011g0032 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
intron_variant | MODIFIER | c.225+652C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756465 | ||||||
| chr1:12756471
|
C | T | 1 | a0001c0001t0005g0093 | 1 | NA18947.hp2 | intron_variant | MODIFIER | c.225+658C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756471 | ||||||
| chr1:12756516
|
A | G | 2 | a0001c0001t0001g0009a0001c0001t0001g0015 | 7 | HG00639.hp2 HG00738.hp1 HG00741.hp1 others(4): Show |
intron_variant | MODIFIER | c.225+703A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756516 | ||||||
| chr1:12756673
|
G | T | 1 | a0001c0001t0001g0085 | 1 | HG02922.hp1 | intron_variant | MODIFIER | c.225+860G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756673 | ||||||
| chr1:12756786
|
C | A | 1 | a0001c0001t0002g0078 | 1 | HG01081.hp2 | intron_variant | MODIFIER | c.225+973C>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756786 | ||||||
| chr1:12756825
|
T | A | 1 | a0001c0001t0002g0079 | 1 | NA18747.hp2 | intron_variant | MODIFIER | c.225+1012T>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756825 | ||||||
| chr1:12756827
|
T | G | 64 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(61): Show | 143 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(140): Show |
intron_variant | MODIFIER | c.225+1014T>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756827 | ||||||
| chr1:12756919
|
G | T | 1 | a0001c0001t0045g0096 | 1 | HG02559.hp1 | intron_variant | MODIFIER | c.225+1106G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12756919 | ||||||
| chr1:12757007
|
A | G | 2 | a0001c0001t0002g0078a0001c0001t0002g0084 | 2 | HG00099.hp1 HG01081.hp2 |
intron_variant | MODIFIER | c.225+1194A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757007 | ||||||
| chr1:12757112
|
C | T | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.225+1299C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757112 | ||||||
| chr1:12757136
|
T | C | 8 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(5): Show | 19 | HG01106.hp2 HG01192.hp1 HG01243.hp1 others(16): Show |
intron_variant | MODIFIER | c.225+1323T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757136 | ||||||
| chr1:12757271
|
C | CG | 4 | a0001c0001t0007g0091a0001c0001t0013g0092a0002c0002t0003g0101others(1): Show | 4 | HG00423.hp1 HG03927.hp1 NA18967.hp1 others(1): Show |
intron_variant | MODIFIER | c.225+1458_225+1459i others(3): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757271 | ||||||
| chr1:12757272
|
A | C | 4 | a0001c0001t0007g0091a0001c0001t0013g0092a0002c0002t0003g0101others(1): Show | 4 | HG00423.hp1 HG03927.hp1 NA18967.hp1 others(1): Show |
intron_variant | MODIFIER | c.225+1459A>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757272 | ||||||
| chr1:12757272
|
A | G | 49 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(46): Show | 117 | HG00323.hp1 HG00323.hp2 HG00438.hp1 others(114): Show |
intron_variant | MODIFIER | c.225+1459A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757272 | ||||||
| chr1:12757302
|
A | G | 1 | a0001c0008t0001g0070 | 1 | HG03942.hp1 | intron_variant | MODIFIER | c.225+1489A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757302 | ||||||
| chr1:12757305
|
C | T | 1 | a0001c0001t0002g0083 | 1 | HG01952.hp2 | intron_variant | MODIFIER | c.225+1492C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757305 | ||||||
| chr1:12757328
|
TTCTTTTC others(10): Show |
T | 50 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(47): Show | 118 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(115): Show |
intron_variant | MODIFIER | c.225+1531_225+1547d others(19): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | INFO_REALIGN_3_PRIME | chr1 | 12757328 | |||||
| chr1:12757333
|
TTCTTTTC others(5): Show |
T | 1 | a0001c0001t0044g0109 | 1 | HG02451.hp1 | intron_variant | MODIFIER | c.225+1534_225+1545d others(14): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | INFO_REALIGN_3_PRIME | chr1 | 12757333 | |||||
| chr1:12757340
|
CTTTTCTC others(1): Show |
C | 3 | a0002c0002t0003g0098a0002c0002t0003g0104a0003c0005t0023g0035 | 3 | HG01175.hp2 NA19060.hp1 NA19240.hp2 |
intron_variant | MODIFIER | c.225+1531_225+1538d others(10): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | INFO_REALIGN_3_PRIME | chr1 | 12757340 | |||||
| chr1:12757354
|
TTTC | T | 3 | a0002c0002t0003g0098a0002c0002t0003g0104a0003c0005t0023g0035 | 3 | HG01175.hp2 NA19060.hp1 NA19240.hp2 |
intron_variant | MODIFIER | c.225+1544_225+1546d others(5): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | INFO_REALIGN_3_PRIME | chr1 | 12757354 | |||||
| chr1:12757367
|
C | CT | 36 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0019g0117others(33): Show | 85 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(82): Show |
intron_variant | MODIFIER | c.225+1562dupT | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | INFO_REALIGN_3_PRIME | chr1 | 12757367 | |||||
| chr1:12757379
|
T | G | 2 | a0005c0003t0011g0018a0005c0003t0011g0032 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
intron_variant | MODIFIER | c.225+1566T>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757379 | ||||||
| chr1:12757390
|
CT | C | 62 | a0001c0001t0004g0005a0001c0001t0004g0040a0001c0001t0004g0046others(59): Show | 141 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(138): Show |
intron_variant | MODIFIER | c.225+1586delT | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | INFO_REALIGN_3_PRIME | chr1 | 12757390 | |||||
| chr1:12757405
|
A | G | 1 | a0002c0002t0003g0105 | 1 | HG02723.hp1 | intron_variant | MODIFIER | c.225+1592A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757405 | ||||||
| chr1:12757567
|
G | A | 5 | a0001c0001t0001g0080a0001c0001t0002g0022a0001c0001t0002g0078others(2): Show | 5 | HG00099.hp1 HG01081.hp1 HG01081.hp2 others(2): Show |
intron_variant | MODIFIER | c.226-1743G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757567 | ||||||
| chr1:12757724
|
G | A | 1 | a0001c0001t0026g0108 | 1 | HG02717.hp1 | intron_variant | MODIFIER | c.226-1586G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757724 | ||||||
| chr1:12757736
|
G | C | 12 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(9): Show | 28 | HG00597.hp2 HG01516.hp1 HG01517.hp1 others(25): Show |
intron_variant | MODIFIER | c.226-1574G>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757736 | ||||||
| chr1:12757768
|
T | C | 1 | a0001c0001t0020g0114 | 1 | HG03195.hp1 | intron_variant | MODIFIER | c.226-1542T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757768 | ||||||
| chr1:12757793
|
T | C | 1 | a0001c0001t0048g0111 | 1 | HG02559.hp2 | intron_variant | MODIFIER | c.226-1517T>C | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757793 | ||||||
| chr1:12757837
|
C | T | 1 | a0001c0001t0001g0069 | 1 | HG03130.hp1 | intron_variant | MODIFIER | c.226-1473C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757837 | ||||||
| chr1:12757842
|
C | T | 1 | a0003c0005t0022g0034 | 1 | HG00642.hp1 | intron_variant | MODIFIER | c.226-1468C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757842 | ||||||
| chr1:12757944
|
C | T | 1 | a0001c0001t0001g0052 | 1 | HG00741.hp2 | intron_variant | MODIFIER | c.226-1366C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12757944 | ||||||
| chr1:12758012
|
A | G | 52 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(49): Show | 120 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(117): Show |
intron_variant | MODIFIER | c.226-1298A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12758012 | ||||||
| chr1:12758053
|
G | T | 1 | a0001c0001t0044g0109 | 1 | HG02451.hp1 | intron_variant | MODIFIER | c.226-1257G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12758053 | ||||||
| chr1:12758206
|
GTAAC | G | 13 | a0001c0001t0001g0007a0001c0001t0004g0005a0001c0001t0004g0040others(10): Show | 27 | HG01106.hp2 HG01167.hp2 HG01192.hp1 others(24): Show |
intron_variant | MODIFIER | c.226-1103_226-1100d others(6): Show |
CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12758206 | ||||||
| chr1:12758553
|
C | T | 3 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0021g0116 | 5 | HG01109.hp1 HG02280.hp2 HG02717.hp2 others(2): Show |
intron_variant | MODIFIER | c.226-757C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12758553 | ||||||
| chr1:12758908
|
A | G | 14 | a0001c0001t0001g0007a0001c0001t0004g0005a0001c0001t0004g0040others(11): Show | 30 | HG01106.hp2 HG01167.hp2 HG01192.hp1 others(27): Show |
intron_variant | MODIFIER | c.226-402A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12758908 | ||||||
| chr1:12759130
|
T | A | 66 | a0001c0001t0001g0007a0001c0001t0004g0005a0001c0001t0004g0040others(63): Show | 148 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(145): Show |
intron_variant | MODIFIER | c.226-180T>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12759130 | ||||||
| chr1:12759228
|
A | G | 3 | a0001c0001t0012g0019a0001c0001t0012g0115a0001c0001t0021g0116 | 5 | HG01109.hp1 HG02280.hp2 HG02717.hp2 others(2): Show |
intron_variant | MODIFIER | c.226-82A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12759228 | ||||||
| chr1:12759284
|
C | T | 2 | a0001c0001t0008g0081a0001c0001t0026g0108 | 2 | HG02717.hp1 NA21309.hp2 |
intron_variant | MODIFIER | c.226-26C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 2/3 | chr1 | 12759284 | ||||||
| chr1:12759607
|
C | T | 1 | a0001c0001t0029g0045 | 1 | HG03195.hp2 | intron_variant | MODIFIER | c.403+120C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12759607 | ||||||
| chr1:12759761
|
C | T | 1 | a0001c0001t0020g0114 | 1 | HG03195.hp1 | intron_variant | MODIFIER | c.403+274C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12759761 | ||||||
| chr1:12759969
|
A | T | 1 | a0001c0001t0045g0096 | 1 | HG02559.hp1 | intron_variant | MODIFIER | c.403+482A>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12759969 | ||||||
| chr1:12760000
|
A | G | 1 | a0001c0001t0001g0061 | 1 | HG04184.hp2 | intron_variant | MODIFIER | c.403+513A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760000 | ||||||
| chr1:12760132
|
A | G | 1 | a0001c0001t0006g0082 | 1 | NA18954.hp1 | intron_variant | MODIFIER | c.404-638A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760132 | ||||||
| chr1:12760264
|
C | G | 2 | a0005c0003t0011g0018a0005c0003t0011g0032 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
intron_variant | MODIFIER | c.404-506C>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760264 | ||||||
| chr1:12760350
|
A | G | 2 | a0005c0003t0011g0018a0005c0003t0011g0032 | 5 | HG02055.hp2 HG02647.hp1 HG03453.hp1 others(2): Show |
intron_variant | MODIFIER | c.404-420A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760350 | ||||||
| chr1:12760501
|
A | G | 49 | a0001c0001t0005g0004a0001c0001t0005g0027a0001c0001t0005g0093others(46): Show | 113 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(110): Show |
intron_variant | MODIFIER | c.404-269A>G | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760501 | ||||||
| chr1:12760558
|
G | T | 25 | a0002c0002t0003g0003a0002c0002t0003g0008a0002c0002t0003g0017others(22): Show | 68 | HG00323.hp1 HG00323.hp2 HG00423.hp1 others(65): Show |
intron_variant | MODIFIER | c.404-212G>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760558 | ||||||
| chr1:12760697
|
C | T | 1 | a0001c0001t0039g0051 | 1 | NA19043.hp1 | intron_variant | MODIFIER | c.404-73C>T | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760697 | ||||||
| chr1:12760698
|
G | A | 6 | a0001c0001t0019g0117a0001c0001t0021g0116a0004c0004t0010g0020others(3): Show | 8 | HG02257.hp1 HG02717.hp2 HG02896.hp2 others(5): Show |
intron_variant | MODIFIER | c.404-72G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760698 | ||||||
| chr1:12760708
|
G | A | 6 | a0001c0001t0019g0117a0001c0001t0021g0116a0004c0004t0010g0020others(3): Show | 8 | HG02257.hp1 HG02717.hp2 HG02896.hp2 others(5): Show |
intron_variant | MODIFIER | c.404-62G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760708 | ||||||
| chr1:12760732
|
G | A | 1 | a0001c0001t0013g0092 | 1 | HG03927.hp1 | intron_variant | MODIFIER | c.404-38G>A | CFAP107 | ENSG00000157330.10 | transcript | ENST00000614859.5 | protein_coding | 3/3 | chr1 | 12760732 |
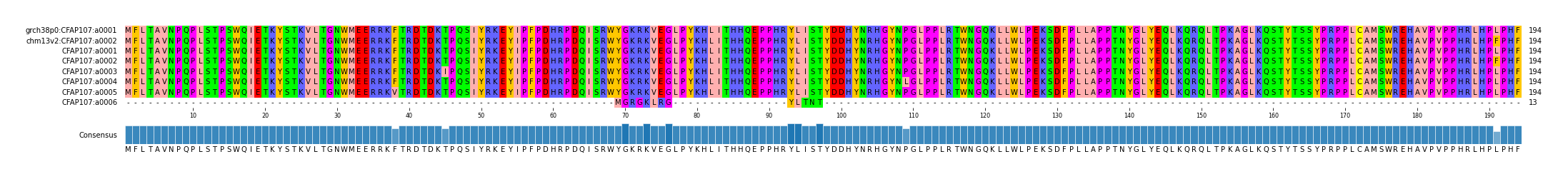 Multiple Alignments (by aa haplotype frequency)
Multiple Alignments (by aa haplotype frequency)
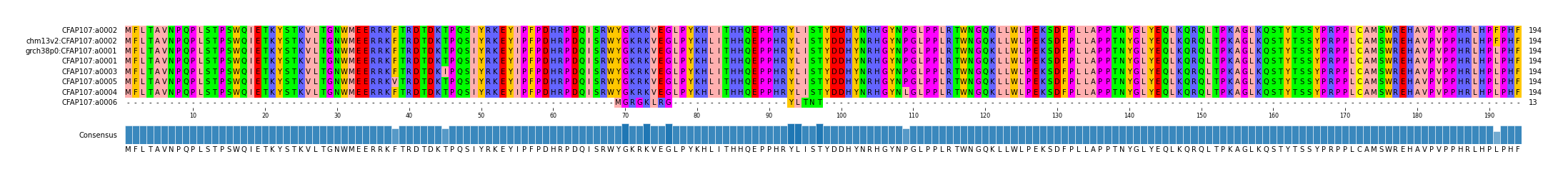 Multiple Alignments (by aa haplotype similarity)
Multiple Alignments (by aa haplotype similarity)