Search by Genic region
| geneid | 91689 |
|---|---|
| ensemblid | ENSG00000183172.9 |
| hgncid | 25055 |
| symbol | SMDT1 |
| name | single-pass membrane protein with aspartate rich tail 1 |
| refseq_nuc | NM_033318.5 |
| refseq_prot | NP_201575.3 |
| ensembl_nuc | ENST00000331479.4 |
| ensembl_prot | ENSP00000327467.3 |
| mane_status | MANE Select |
| chr | chr22 |
| start | 42079700 |
| end | 42084284 |
| strand | + |
| ver | v1.2 |
| region | chr22:42079700-42084284 |
| region5000 | chr22:42074700-42089284 |
| regionname0 | SMDT1_chr22_42079700_42084284 |
| regionname5000 | SMDT1_chr22_42074700_42089284 |
| ahapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
alen | total | AFR | AMR | EAS | EUR | SAS | JPT | regionname | genename | aa | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001 | 0/1 | 107 | 392 | 94 | 74 | 163 | 16 | 44 | 131 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0002 | 0/0 | 107 | 1 | 0 | 0 | 1 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| chapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
clen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | cseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| c0001 | 0/1 | 324 | 392 | 94 | 74 | 163 | 16 | 44 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| c0002 | 0/0 | 324 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| thapid | grch38chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
tlen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | tseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| t0001 | 0/1 | 1239 | 367 | 80 | 74 | 162 | 15 | 35 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0002 | 0/0 | 1239 | 11 | 11 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0003 | 0/0 | 1239 | 5 | 0 | 0 | 0 | 0 | 5 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0004 | 0/0 | 1239 | 3 | 0 | 0 | 0 | 0 | 3 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0005 | 0/0 | 1239 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0006 | 0/0 | 1239 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0007 | 0/0 | 1239 | 1 | 0 | 0 | 0 | 1 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0008 | 0/0 | 1229 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0009 | 0/0 | 1239 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| t0010 | 0/0 | 1239 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| ghapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
total | AFR | AMR | EAS | EUR | SAS | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| g0001 | 0/0 | 229 | 28 | 43 | 126 | 9 | 23 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0002 | 0/1 | 91 | 10 | 24 | 32 | 7 | 17 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0003 | 0/0 | 24 | 22 | 2 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0004 | 0/0 | 17 | 16 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0005 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0006 | 0/0 | 3 | 0 | 1 | 1 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0007 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0008 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0009 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0010 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0011 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0012 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0013 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0014 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0015 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0016 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0017 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0018 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0019 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0020 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0021 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0022 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0023 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0024 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0025 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| g0026 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| achapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
clen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | cseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001 | 0/1 | 324 | 392 | 94 | 74 | 163 | 16 | 44 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0002c0002 | 0/0 | 324 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| acthapid | grch38chm13v2 | tlen | total | AFR | AMR | EAS | EUR | SAS | regionname | genename | tseq | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001t0001 | 0/1 | 1562 | 366 | 80 | 74 | 161 | 15 | 35 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0002 | 0/0 | 1562 | 11 | 11 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0003 | 0/0 | 1562 | 5 | 0 | 0 | 0 | 0 | 5 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0004 | 0/0 | 1562 | 3 | 0 | 0 | 0 | 0 | 3 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0005 | 0/0 | 1562 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0006 | 0/0 | 1562 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0007 | 0/0 | 1562 | 1 | 0 | 0 | 0 | 1 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0008 | 0/0 | 1552 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0009 | 0/0 | 1562 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0001c0001t0010 | 0/0 | 1562 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| a0002c0002t0001 | 0/0 | 1562 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | copy fasta | chr22 | 42074700 | 42089284 |
| actghapid | grch38/chm13v2 1/0: The haplotype type is the same as GRCh380/1: The haplotype type is the same as CHM13v20/0: The haplotype type matches neither GRCh38 nor CHM13v21/1: The haplotype type is the same on both GRCh38 and CHM13v2
|
total | AFR | AMR | EAS | EUR | SAS | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a0001c0001t0001g0001 | 0/0 | 215 | 17 | 43 | 124 | 8 | 23 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0002 | 0/1 | 81 | 10 | 24 | 31 | 7 | 8 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0003 | 0/0 | 24 | 22 | 2 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0004 | 0/0 | 17 | 16 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0005 | 0/0 | 3 | 3 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0006 | 0/0 | 3 | 0 | 1 | 1 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0007 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0008 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0009 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0010 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0011 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0012 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0013 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0014 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0015 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0016 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0017 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0019 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0020 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0021 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0022 | 0/0 | 1 | 0 | 1 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0023 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0024 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0025 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0001g0026 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0002g0001 | 0/0 | 10 | 10 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0002g0018 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0003g0002 | 0/0 | 5 | 0 | 0 | 0 | 0 | 5 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0004g0002 | 0/0 | 3 | 0 | 0 | 0 | 0 | 3 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0005g0007 | 0/0 | 2 | 2 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0006g0001 | 0/0 | 1 | 1 | 0 | 0 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0007g0001 | 0/0 | 1 | 0 | 0 | 0 | 1 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0008g0001 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0009g0002 | 0/0 | 1 | 0 | 0 | 0 | 0 | 1 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0001c0001t0010g0001 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| a0002c0002t0001g0002 | 0/0 | 1 | 0 | 0 | 1 | 0 | 0 | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| sampleid | ID haplotypeid
|
ahapid | chapid | thapid | ghapid | gpopname | popname | regionname | genename | chr | start | end |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| HG00099 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | GBR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00099 | hp2 | a0001 | c0001 | t0001 | g0002 | EUR | GBR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00140 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | GBR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00140 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | GBR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00280 | hp1 | a0001 | c0001 | t0001 | g0002 | EUR | FIN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00280 | hp2 | a0001 | c0001 | t0001 | g0002 | EUR | FIN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00323 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | FIN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00323 | hp2 | a0001 | c0001 | t0001 | g0002 | EUR | FIN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00423 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00423 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00438 | hp1 | a0001 | c0001 | t0008 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00438 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00544 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00544 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00558 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00558 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00621 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00621 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00639 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00639 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00642 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00642 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00673 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00673 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00733 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00733 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00735 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00735 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00741 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG00741 | hp2 | a0001 | c0001 | t0001 | g0015 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01069 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01069 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01071 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01071 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01074 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01074 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01081 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01081 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01099 | hp1 | a0001 | c0001 | t0001 | g0022 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01099 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01106 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01106 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01109 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01109 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01167 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01167 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01168 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01168 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01175 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01175 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01192 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01192 | hp2 | a0001 | c0001 | t0001 | g0017 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01243 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01243 | hp2 | a0001 | c0001 | t0001 | g0003 | AMR | PUR | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01255 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01255 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01256 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01256 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01258 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01258 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01261 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01261 | hp2 | a0001 | c0001 | t0001 | g0006 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01346 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01346 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01358 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01358 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01361 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01361 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01433 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01433 | hp2 | a0001 | c0001 | t0001 | g0003 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01515 | hp1 | a0001 | c0001 | t0007 | g0001 | EUR | IBS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01515 | hp2 | a0001 | c0001 | t0001 | g0002 | EUR | IBS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01516 | hp1 | a0001 | c0001 | t0001 | g0002 | EUR | IBS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01516 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | IBS | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01884 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01884 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01891 | hp1 | a0001 | c0001 | t0002 | g0018 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01891 | hp2 | a0001 | c0001 | t0001 | g0005 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01928 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01928 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01934 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01934 | hp2 | a0001 | c0001 | t0001 | g0004 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01943 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01943 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01952 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01952 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01978 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01978 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01981 | hp1 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01981 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01993 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01993 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02004 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02004 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02027 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02027 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02055 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02055 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02071 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02071 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02074 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02074 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02080 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02080 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02083 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02083 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02129 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02129 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02145 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02145 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02148 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02148 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02155 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CDX | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02155 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CDX | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02165 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CDX | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02165 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CDX | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02257 | hp1 | a0001 | c0001 | t0001 | g0005 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02257 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02258 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02258 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02273 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02273 | hp2 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02280 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02280 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02293 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02293 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | PEL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02451 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02451 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02523 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02523 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | KHV | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02572 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02572 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02602 | hp1 | a0001 | c0001 | t0003 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02602 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02615 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02615 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02622 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02622 | hp2 | a0001 | c0001 | t0001 | g0009 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02630 | hp1 | a0001 | c0001 | t0001 | g0009 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02630 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02647 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02647 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02683 | hp1 | a0001 | c0001 | t0001 | g0025 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02683 | hp2 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02698 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02698 | hp2 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02717 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02717 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02723 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02723 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02735 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02735 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02738 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02738 | hp2 | a0001 | c0001 | t0001 | g0006 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02818 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02818 | hp2 | a0001 | c0001 | t0001 | g0007 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02886 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02886 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02895 | hp1 | a0001 | c0001 | t0002 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02895 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02896 | hp1 | a0001 | c0001 | t0001 | g0008 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02896 | hp2 | a0001 | c0001 | t0001 | g0024 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02897 | hp1 | a0001 | c0001 | t0002 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02897 | hp2 | a0001 | c0001 | t0001 | g0008 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02965 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02965 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02970 | hp1 | a0001 | c0001 | t0001 | g0010 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02970 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02976 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02976 | hp2 | a0001 | c0001 | t0001 | g0011 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03017 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03017 | hp2 | a0001 | c0001 | t0009 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03041 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03041 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03098 | hp1 | a0001 | c0001 | t0002 | g0001 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03098 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03130 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03130 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03139 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03139 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03195 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03195 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03225 | hp1 | a0001 | c0001 | t0006 | g0001 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03225 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03453 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03453 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03486 | hp1 | a0001 | c0001 | t0001 | g0026 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03486 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03490 | hp1 | a0001 | c0001 | t0004 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03490 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03491 | hp1 | a0001 | c0001 | t0003 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03491 | hp2 | a0001 | c0001 | t0004 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03492 | hp1 | a0001 | c0001 | t0003 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03492 | hp2 | a0001 | c0001 | t0004 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03516 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03516 | hp2 | a0001 | c0001 | t0001 | g0023 | AFR | ESN | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03540 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03540 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | GWD | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03579 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03579 | hp2 | a0001 | c0001 | t0005 | g0007 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03654 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03654 | hp2 | a0001 | c0001 | t0003 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03669 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03669 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03704 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03704 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03710 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03710 | hp2 | a0001 | c0001 | t0003 | g0002 | SAS | PJL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03831 | hp1 | a0001 | c0001 | t0001 | g0012 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03831 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03834 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03834 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03927 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03927 | hp2 | a0001 | c0001 | t0001 | g0002 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03942 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03942 | hp2 | a0001 | c0001 | t0001 | g0002 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04115 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04115 | hp2 | a0001 | c0001 | t0001 | g0019 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04184 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04184 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | BEB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04199 | hp1 | a0001 | c0001 | t0001 | g0002 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04199 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04204 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04204 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04228 | hp1 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG04228 | hp2 | a0001 | c0001 | t0001 | g0001 | SAS | STU | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18522 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | YRI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18522 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | YRI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18747 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | CHB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18747 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | CHB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18906 | hp1 | a0001 | c0001 | t0001 | g0005 | AFR | YRI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18906 | hp2 | a0001 | c0001 | t0001 | g0004 | AFR | YRI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18939 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18939 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18940 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18940 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18941 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18941 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18945 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18945 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18947 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18947 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18949 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18949 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18950 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18950 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18951 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18951 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18953 | hp1 | a0001 | c0001 | t0001 | g0016 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18953 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18957 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18957 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18959 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18959 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18961 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18961 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18962 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18962 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18963 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18963 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18964 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18964 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18965 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18965 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18966 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18966 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18967 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18967 | hp2 | a0001 | c0001 | t0001 | g0014 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18968 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18968 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18969 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18969 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18971 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18971 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18972 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18972 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18973 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18973 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18974 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18974 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18975 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18975 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18978 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18978 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18979 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18979 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18980 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18980 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18981 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18981 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18982 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18982 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18984 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18984 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18986 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18986 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18987 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18987 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18989 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18989 | hp2 | a0001 | c0001 | t0001 | g0006 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18990 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18990 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18991 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18991 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18992 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18992 | hp2 | a0001 | c0001 | t0010 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18994 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18994 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18998 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18998 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18999 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA18999 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19000 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19000 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19001 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19001 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19004 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19004 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19010 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19010 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19011 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19011 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19030 | hp1 | a0001 | c0001 | t0001 | g0004 | AFR | LWK | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19030 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | LWK | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19043 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | LWK | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19043 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | LWK | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19054 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19054 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19055 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19055 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19057 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19057 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19060 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19060 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19065 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19065 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19068 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19068 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19070 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19070 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19074 | hp1 | a0002 | c0002 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19074 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19075 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19075 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19077 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19077 | hp2 | a0001 | c0001 | t0001 | g0020 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19080 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19080 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19081 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19081 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19083 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19083 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19084 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19084 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19085 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19085 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19086 | hp1 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19086 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19087 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19087 | hp2 | a0001 | c0001 | t0001 | g0002 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19088 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19088 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19089 | hp1 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19089 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19090 | hp1 | a0001 | c0001 | t0001 | g0013 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19090 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19091 | hp1 | a0001 | c0001 | t0001 | g0021 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19091 | hp2 | a0001 | c0001 | t0001 | g0001 | EAS | JPT | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19240 | hp1 | a0001 | c0001 | t0001 | g0011 | AFR | YRI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA19240 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | YRI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20129 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | ASW | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20129 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | ASW | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20752 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | TSI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20752 | hp2 | a0001 | c0001 | t0001 | g0001 | EUR | TSI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20805 | hp1 | a0001 | c0001 | t0001 | g0001 | EUR | TSI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20805 | hp2 | a0001 | c0001 | t0001 | g0002 | EUR | TSI | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01123 | hp1 | a0001 | c0001 | t0001 | g0001 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG01123 | hp2 | a0001 | c0001 | t0001 | g0002 | AMR | CLM | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02109 | hp1 | a0001 | c0001 | t0001 | g0010 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02109 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02486 | hp1 | a0001 | c0001 | t0005 | g0007 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02486 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02559 | hp1 | a0001 | c0001 | t0001 | g0002 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG02559 | hp2 | a0001 | c0001 | t0001 | g0003 | AFR | ACB | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03471 | hp1 | a0001 | c0001 | t0002 | g0001 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG03471 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | MSL | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG06807 | hp1 | a0001 | c0001 | t0001 | g0001 | AFR | USA | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| HG06807 | hp2 | a0001 | c0001 | t0001 | g0002 | AFR | USA | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20300 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | USA | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA20300 | hp2 | a0001 | c0001 | t0001 | g0001 | AFR | USA | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA21309 | hp1 | a0001 | c0001 | t0001 | g0003 | AFR | LWK | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| NA21309 | hp2 | a0001 | c0001 | t0002 | g0001 | AFR | LWK | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| homoSapiens_chm13v2 | hp1 | a0001 | c0001 | t0001 | g0002 | REF | REF | SMDT1_chr22_42074700_42089284 | SMDT1 | chr22 | 42074700 | 42089284 |
| chr:pos | ref | alt | # # of ahapid:amino-acid(protein) level |
ahapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr22:42082041
|
G | C | 1 | a0002 | 1 | NA19074.hp1 | missense_variant | MODERATE | c.303G>C | p.Glu101Asp | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 2/3 | 372/1562 | 303/324 | 101/107 | chr22 | 42082041 |
| chr:pos | ref | alt | # # of chapid |
chapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|
| chr:pos | ref | alt | # # of thapid:transcript level |
thapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | transcript_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr22:42083232
|
A | T | 1 | a0001c0001t0010 | 1 | NA18992.hp2 | 3_prime_UTR_variant | MODIFIER | c.*117A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1170 | chr22 | 42083232 | |||||
| chr22:42083341
|
A | C | 1 | a0001c0001t0009 | 1 | HG03017.hp2 | 3_prime_UTR_variant | MODIFIER | c.*226A>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1279 | chr22 | 42083341 | |||||
| chr22:42083361
|
G | A | 1 | a0001c0001t0005 | 2 | HG02486.hp1 HG03579.hp2 |
3_prime_UTR_variant | MODIFIER | c.*246G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1299 | chr22 | 42083361 | |||||
| chr22:42083721
|
A | G | 1 | a0001c0001t0003 | 5 | HG02602.hp1 HG03491.hp1 HG03492.hp1 others(2): Show |
3_prime_UTR_variant | MODIFIER | c.*606A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1659 | chr22 | 42083721 | |||||
| chr22:42083823
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*708C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1761 | chr22 | 42083823 | |||||
| chr22:42083825
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*710C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1763 | chr22 | 42083825 | |||||
| chr22:42083827
|
T | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*712T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1765 | chr22 | 42083827 | |||||
| chr22:42083829
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*714C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1767 | chr22 | 42083829 | |||||
| chr22:42083831
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*716G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1769 | chr22 | 42083831 | |||||
| chr22:42083838
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*723C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1776 | chr22 | 42083838 | |||||
| chr22:42083846
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*731G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1784 | chr22 | 42083846 | |||||
| chr22:42083858
|
A | C | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*743A>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1796 | chr22 | 42083858 | |||||
| chr22:42083859
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*744A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1797 | chr22 | 42083859 | |||||
| chr22:42083860
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*745A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1798 | chr22 | 42083860 | |||||
| chr22:42083861
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*746C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1799 | chr22 | 42083861 | |||||
| chr22:42083864
|
A | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*749A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1802 | chr22 | 42083864 | |||||
| chr22:42083865
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*750A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1803 | chr22 | 42083865 | |||||
| chr22:42083866
|
A | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*751A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1804 | chr22 | 42083866 | |||||
| chr22:42083867
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*752G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1805 | chr22 | 42083867 | |||||
| chr22:42083877
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*762C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1815 | chr22 | 42083877 | |||||
| chr22:42083881
|
T | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*766T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1819 | chr22 | 42083881 | |||||
| chr22:42083882
|
G | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*767G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1820 | chr22 | 42083882 | |||||
| chr22:42083883
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*768C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1821 | chr22 | 42083883 | |||||
| chr22:42083885
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*770C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1823 | chr22 | 42083885 | |||||
| chr22:42083889
|
T | C | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*774T>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1827 | chr22 | 42083889 | |||||
| chr22:42083891
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*776C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1829 | chr22 | 42083891 | |||||
| chr22:42083892
|
C | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*777C>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1830 | chr22 | 42083892 | |||||
| chr22:42083894
|
G | A | 1 | a0001c0001t0002 | 11 | HG01891.hp1 HG02486.hp2 HG02895.hp1 others(8): Show |
3_prime_UTR_variant | MODIFIER | c.*779G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1832 | chr22 | 42083894 | |||||
| chr22:42083903
|
A | C | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*788A>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1841 | chr22 | 42083903 | |||||
| chr22:42083904
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*789A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1842 | chr22 | 42083904 | |||||
| chr22:42083906
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*791A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1844 | chr22 | 42083906 | |||||
| chr22:42083908
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*793G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1846 | chr22 | 42083908 | |||||
| chr22:42083909
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*794A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1847 | chr22 | 42083909 | |||||
| chr22:42083910
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*795A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1848 | chr22 | 42083910 | |||||
| chr22:42083913
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*798C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1851 | chr22 | 42083913 | |||||
| chr22:42083919
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*804C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1857 | chr22 | 42083919 | |||||
| chr22:42083920
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*805C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1858 | chr22 | 42083920 | |||||
| chr22:42083924
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*809C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1862 | chr22 | 42083924 | |||||
| chr22:42083928
|
T | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*813T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1866 | chr22 | 42083928 | |||||
| chr22:42083931
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*816G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1869 | chr22 | 42083931 | |||||
| chr22:42083942
|
ATGAGACC others(3): Show |
A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*828_*837delTGAGAC others(4): Show |
SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1881 | chr22 | 42083942 | |||||
| chr22:42083963
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*848G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1901 | chr22 | 42083963 | |||||
| chr22:42083966
|
T | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*851T>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1904 | chr22 | 42083966 | |||||
| chr22:42083968
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*853C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1906 | chr22 | 42083968 | |||||
| chr22:42083969
|
T | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*854T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1907 | chr22 | 42083969 | |||||
| chr22:42083974
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*859C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1912 | chr22 | 42083974 | |||||
| chr22:42083975
|
A | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*860A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1913 | chr22 | 42083975 | |||||
| chr22:42083978
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*863C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1916 | chr22 | 42083978 | |||||
| chr22:42083979
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*864C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1917 | chr22 | 42083979 | |||||
| chr22:42083982
|
A | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*867A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1920 | chr22 | 42083982 | |||||
| chr22:42083983
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*868G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1921 | chr22 | 42083983 | |||||
| chr22:42083985
|
G | C | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*870G>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1923 | chr22 | 42083985 | |||||
| chr22:42083986
|
G | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*871G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1924 | chr22 | 42083986 | |||||
| chr22:42083991
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*876A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1929 | chr22 | 42083991 | |||||
| chr22:42083995
|
T | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*880T>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1933 | chr22 | 42083995 | |||||
| chr22:42083997
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*882G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1935 | chr22 | 42083997 | |||||
| chr22:42083998
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*883A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1936 | chr22 | 42083998 | |||||
| chr22:42083999
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*884C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1937 | chr22 | 42083999 | |||||
| chr22:42084003
|
C | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*888C>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1941 | chr22 | 42084003 | |||||
| chr22:42084003
|
C | T | 1 | a0001c0001t0007 | 1 | HG01515.hp1 | 3_prime_UTR_variant | MODIFIER | c.*888C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1941 | chr22 | 42084003 | |||||
| chr22:42084004
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*889C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1942 | chr22 | 42084004 | |||||
| chr22:42084007
|
G | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*892G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1945 | chr22 | 42084007 | |||||
| chr22:42084010
|
G | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*895G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1948 | chr22 | 42084010 | |||||
| chr22:42084012
|
A | C | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*897A>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1950 | chr22 | 42084012 | |||||
| chr22:42084013
|
C | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*898C>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1951 | chr22 | 42084013 | |||||
| chr22:42084016
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*901C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1954 | chr22 | 42084016 | |||||
| chr22:42084017
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*902C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1955 | chr22 | 42084017 | |||||
| chr22:42084018
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*903C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1956 | chr22 | 42084018 | |||||
| chr22:42084019
|
C | A | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*904C>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1957 | chr22 | 42084019 | |||||
| chr22:42084020
|
A | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*905A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1958 | chr22 | 42084020 | |||||
| chr22:42084021
|
C | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*906C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1959 | chr22 | 42084021 | |||||
| chr22:42084026
|
T | G | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*911T>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1964 | chr22 | 42084026 | |||||
| chr22:42084027
|
C | T | 1 | a0001c0001t0008 | 1 | HG00438.hp1 | 3_prime_UTR_variant | MODIFIER | c.*912C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 1965 | chr22 | 42084027 | |||||
| chr22:42084180
|
C | T | 1 | a0001c0001t0004 | 3 | HG03490.hp1 HG03491.hp2 HG03492.hp2 |
3_prime_UTR_variant | MODIFIER | c.*1065C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 2118 | chr22 | 42084180 | |||||
| chr22:42084216
|
A | G | 1 | a0001c0001t0006 | 1 | HG03225.hp1 | 3_prime_UTR_variant | MODIFIER | c.*1101A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 3/3 | 2154 | chr22 | 42084216 |
| chr:pos | ref | alt | # # of ghapid:genebody level |
ghapids | # # of haplotypeids |
haplotypeids | annotation | impact | hgvs_c | hgvs_p | genename | geneid | featuretype | featureid | genebody_biotype | rank | cdna cdna pos length |
cds cds pos length |
aa aa pos length |
distance | status | chr | pos |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chr22:42080172
|
C | T | 1 | a0001c0001t0001g0026 | 1 | HG03486.hp1 | intron_variant | MODIFIER | c.186+218C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080172 | ||||||
| chr22:42080201
|
G | C | 1 | a0001c0001t0001g0012 | 1 | HG03831.hp1 | intron_variant | MODIFIER | c.186+247G>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080201 | ||||||
| chr22:42080246
|
A | G | 1 | a0001c0001t0001g0025 | 1 | HG02683.hp1 | intron_variant | MODIFIER | c.186+292A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080246 | ||||||
| chr22:42080275
|
T | A | 1 | a0001c0001t0001g0013 | 1 | NA19090.hp1 | intron_variant | MODIFIER | c.186+321T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080275 | ||||||
| chr22:42080305
|
T | A | 1 | a0001c0001t0001g0012 | 1 | HG03831.hp1 | intron_variant | MODIFIER | c.186+351T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080305 | ||||||
| chr22:42080305
|
T | C | 1 | a0001c0001t0001g0014 | 1 | NA18967.hp2 | intron_variant | MODIFIER | c.186+351T>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080305 | ||||||
| chr22:42080306
|
G | T | 1 | a0001c0001t0001g0012 | 1 | HG03831.hp1 | intron_variant | MODIFIER | c.186+352G>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080306 | ||||||
| chr22:42080307
|
A | T | 1 | a0001c0001t0001g0012 | 1 | HG03831.hp1 | intron_variant | MODIFIER | c.186+353A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080307 | ||||||
| chr22:42080411
|
G | A | 1 | a0001c0001t0001g0005 | 3 | HG01891.hp2 HG02257.hp1 NA18906.hp1 |
intron_variant | MODIFIER | c.186+457G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080411 | ||||||
| chr22:42080575
|
G | C | 1 | a0001c0001t0001g0015 | 1 | HG00741.hp2 | intron_variant | MODIFIER | c.186+621G>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080575 | ||||||
| chr22:42080684
|
A | G | 4 | a0001c0001t0001g0004a0001c0001t0001g0010a0001c0001t0001g0011others(1): Show | 22 | HG01934.hp2 HG02109.hp1 HG02258.hp2 others(19): Show |
intron_variant | MODIFIER | c.186+730A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080684 | ||||||
| chr22:42080709
|
A | G | 2 | a0001c0001t0001g0007a0001c0001t0005g0007 | 3 | HG02486.hp1 HG02818.hp2 HG03579.hp2 |
intron_variant | MODIFIER | c.186+755A>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080709 | ||||||
| chr22:42080750
|
A | C | 17 | a0001c0001t0001g0002a0001c0001t0001g0004a0001c0001t0001g0005others(14): Show | 127 | HG00099.hp2 HG00280.hp1 HG00280.hp2 others(124): Show |
intron_variant | MODIFIER | c.186+796A>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080750 | ||||||
| chr22:42080766
|
A | T | 16 | a0001c0001t0001g0002a0001c0001t0001g0004a0001c0001t0001g0005others(13): Show | 126 | HG00099.hp2 HG00280.hp1 HG00280.hp2 others(123): Show |
intron_variant | MODIFIER | c.186+812A>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080766 | ||||||
| chr22:42080982
|
G | A | 1 | a0001c0001t0001g0016 | 1 | NA18953.hp1 | intron_variant | MODIFIER | c.187-943G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42080982 | ||||||
| chr22:42081169
|
C | CT | 6 | a0001c0001t0001g0006a0001c0001t0001g0007a0001c0001t0001g0011others(3): Show | 10 | HG01261.hp2 HG02486.hp1 HG02738.hp2 others(7): Show |
intron_variant | MODIFIER | c.187-742dupT | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | INFO_REALIGN_3_PRIME | chr22 | 42081169 | |||||
| chr22:42081251
|
C | G | 3 | a0001c0001t0001g0005a0001c0001t0001g0009a0001c0001t0001g0010 | 7 | HG01891.hp2 HG02109.hp1 HG02257.hp1 others(4): Show |
intron_variant | MODIFIER | c.187-674C>G | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42081251 | ||||||
| chr22:42081333
|
T | C | 1 | a0001c0001t0001g0024 | 1 | HG02896.hp2 | intron_variant | MODIFIER | c.187-592T>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42081333 | ||||||
| chr22:42081383
|
T | A | 1 | a0001c0001t0001g0021 | 1 | NA19091.hp1 | intron_variant | MODIFIER | c.187-542T>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42081383 | ||||||
| chr22:42081461
|
G | A | 1 | a0001c0001t0001g0017 | 1 | HG01192.hp2 | intron_variant | MODIFIER | c.187-464G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42081461 | ||||||
| chr22:42081586
|
G | A | 2 | a0001c0001t0001g0007a0001c0001t0005g0007 | 3 | HG02486.hp1 HG02818.hp2 HG03579.hp2 |
intron_variant | MODIFIER | c.187-339G>A | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42081586 | ||||||
| chr22:42081646
|
G | C | 10 | a0001c0001t0001g0002a0001c0001t0001g0006a0001c0001t0001g0008others(7): Show | 99 | HG00099.hp2 HG00280.hp1 HG00280.hp2 others(96): Show |
intron_variant | MODIFIER | c.187-279G>C | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 1/2 | chr22 | 42081646 | ||||||
| chr22:42082273
|
C | T | 1 | a0001c0001t0002g0018 | 1 | HG01891.hp1 | intron_variant | MODIFIER | c.*3+208C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 2/2 | chr22 | 42082273 | ||||||
| chr22:42082714
|
C | T | 2 | a0001c0001t0001g0003a0001c0001t0001g0026 | 25 | HG01243.hp2 HG01433.hp2 HG01884.hp1 others(22): Show |
intron_variant | MODIFIER | c.*4-405C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 2/2 | chr22 | 42082714 | ||||||
| chr22:42082856
|
C | T | 1 | a0001c0001t0001g0008 | 2 | HG02896.hp1 HG02897.hp2 |
intron_variant | MODIFIER | c.*4-263C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 2/2 | chr22 | 42082856 | ||||||
| chr22:42083066
|
C | T | 1 | a0001c0001t0001g0022 | 1 | HG01099.hp1 | intron_variant | MODIFIER | c.*4-53C>T | SMDT1 | ENSG00000183172.9 | transcript | ENST00000331479.4 | protein_coding | 2/2 | chr22 | 42083066 |
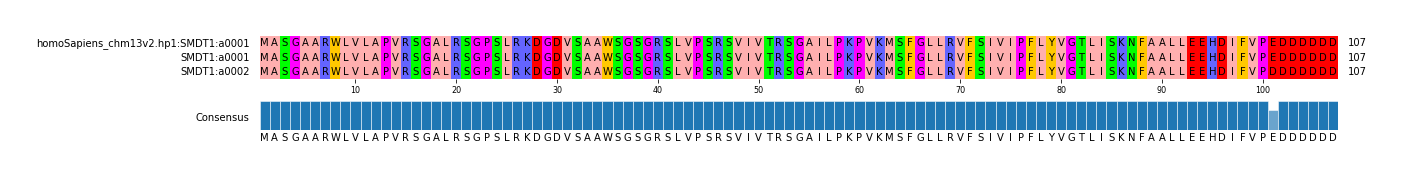 Multiple Alignments (by aa haplotype frequency)
Multiple Alignments (by aa haplotype frequency)
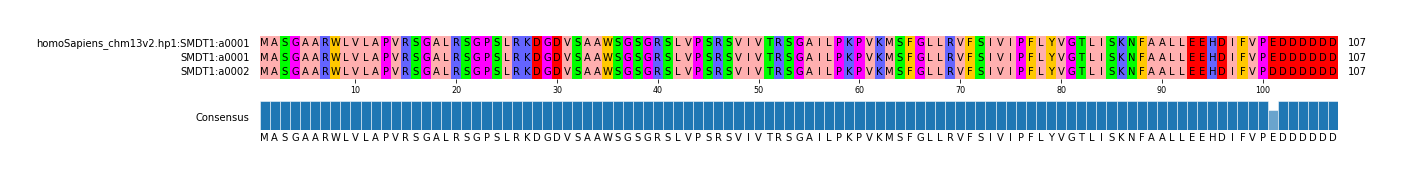 Multiple Alignments (by aa haplotype similarity)
Multiple Alignments (by aa haplotype similarity)